library(tidyverse)Importing and Visualising data
Exploratory data analysis
28 November 2025
Introduction
In this session we will learn how to visualise data using the ggplot2 package.
Importing, data wrangling, and data visualisation constitute a large part of the work you will do in bioinformatics. Today’s session will be a ‘full workflow’.
Learning Objectives
- Learn how to import data in R with the readr package.
- Visualise data with ggplot2.
- Become familiar with the
ggplot()function format. - Use a range of geoms to plot different types of data.
- Practice writing tidy code.
- Practice key skills from day 1 and day 2.
Importing with readr
The readr package is part of the tidyverse.
We can use the read_csv() function to import a csv file into R.
summary() evaluates each column and returns a summary depending on the type of data. For columns of numeric data we see mean, median, min, max and quartile information. For character columns we see the length, and for logical columns we get a count of the number of TRUE and FALSE calls.
summary() is a useful function to understand the shape of your data.
Importing ugly data
We will often encounter data that does not match our expectations for clean and tidy column names.
Go to https://tinyurl.com/uglyDuckling2025 and use the download button to download this example dataset.
Store it in your /data directory.
[1] "Species" "island" "billLengthMM"
[4] "bill_depth_mm" "Flipper_length_mm" "body mass g"
[7] "sex" "Year"
snake_case, camelCase, and spaces??? Who made this object!?
Renaming our ugly data
[1] "species" "island" "bill_length_mm"
[4] "bill_depth_mm" "flipper_length_mm" "body_mass_g"
[7] "sex" "year" What if I had a lot of columns?
The janitor R package has a lot of useful tools for cleaning data.
There are functions and packages for many tasks. Search for existing packages to make your tasks easier.
Importing other data types
Other functions exist for other data:
read_tsv() for tab-delimited files.
read_table() for files where columns are separated by white space.
read_delim() reads in files and guesses what the correct delimiter is. Use in cases where you don’t know what the type of data coming in is.
Data visualisation with ggplot
ggplot (“Grammar of Graphics”): work iteratively, build complexity.
Focus on the format.
Demonstrates a key loop in data analysis: visualise your data, make adjustments, transform the data and visualise again.
The ggplot format
The ggplot format has three parts:
Calling the ggplot function and specifying the data.
Mapping the data: what data is displayed on the x and y axis.
Plotting the data with a geom function - there are different geoms for different types of plots.
- Note that the geom is a new function, which means we need to add a “+” to the previous line.
The ggplot format
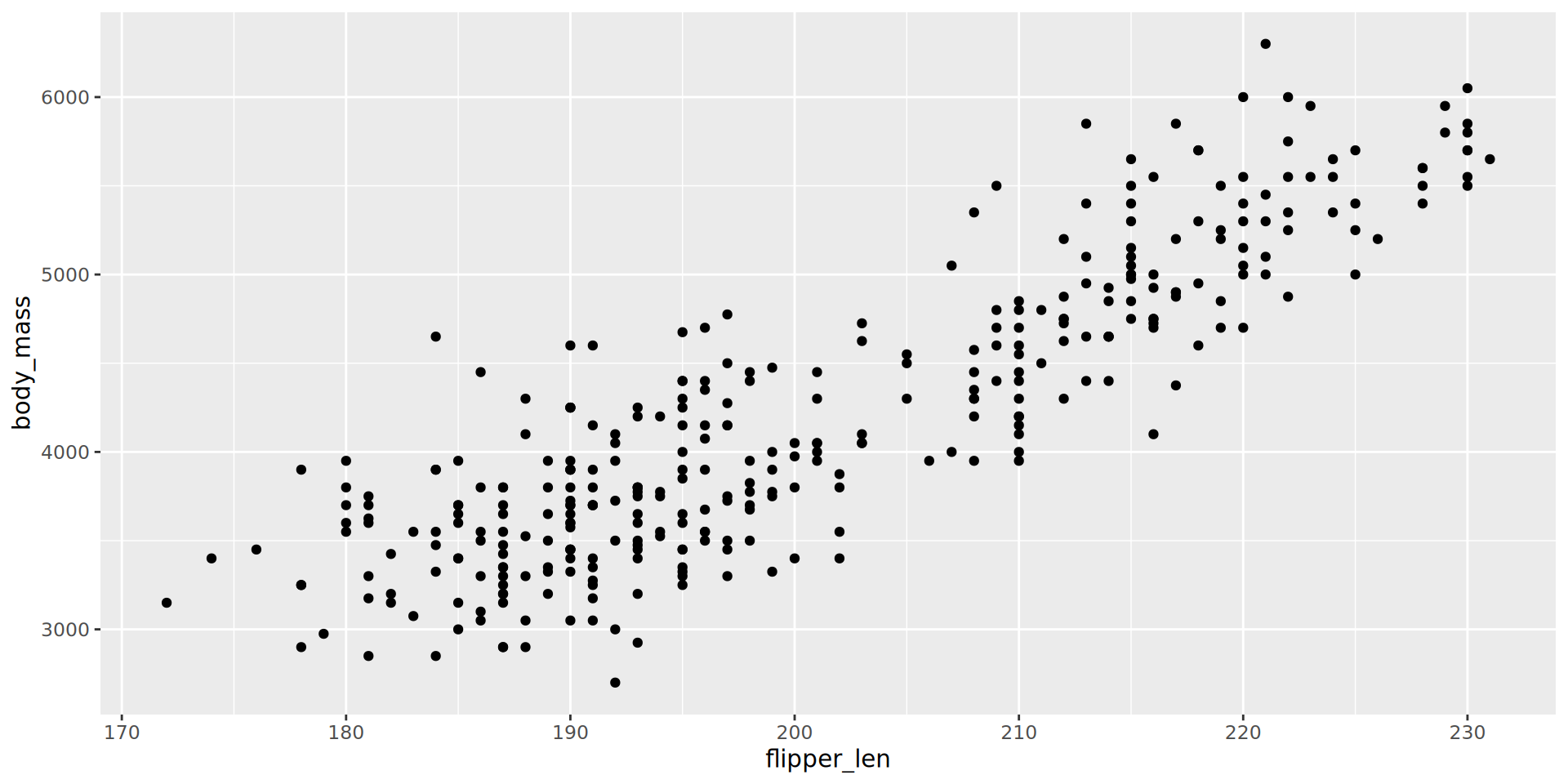
Improving the visualisation
There are many steps we can take to improve our initial visualisation.
Here we will:
Add titles and labels
Improve the visual theme
Re-order the data to reduce cognitive load
Layer multiple geoms
Use geom arguments to change the look of the figure
Titles, labels and theme
Add title and labels with the labs() function.
Themes control the look of the plot window. Try theme_minimal(), theme_dark(), theme_bw(), and many others, for different looks.
- Each new function is preceeded by a “+” on the previous line.
Titles, labels and theme
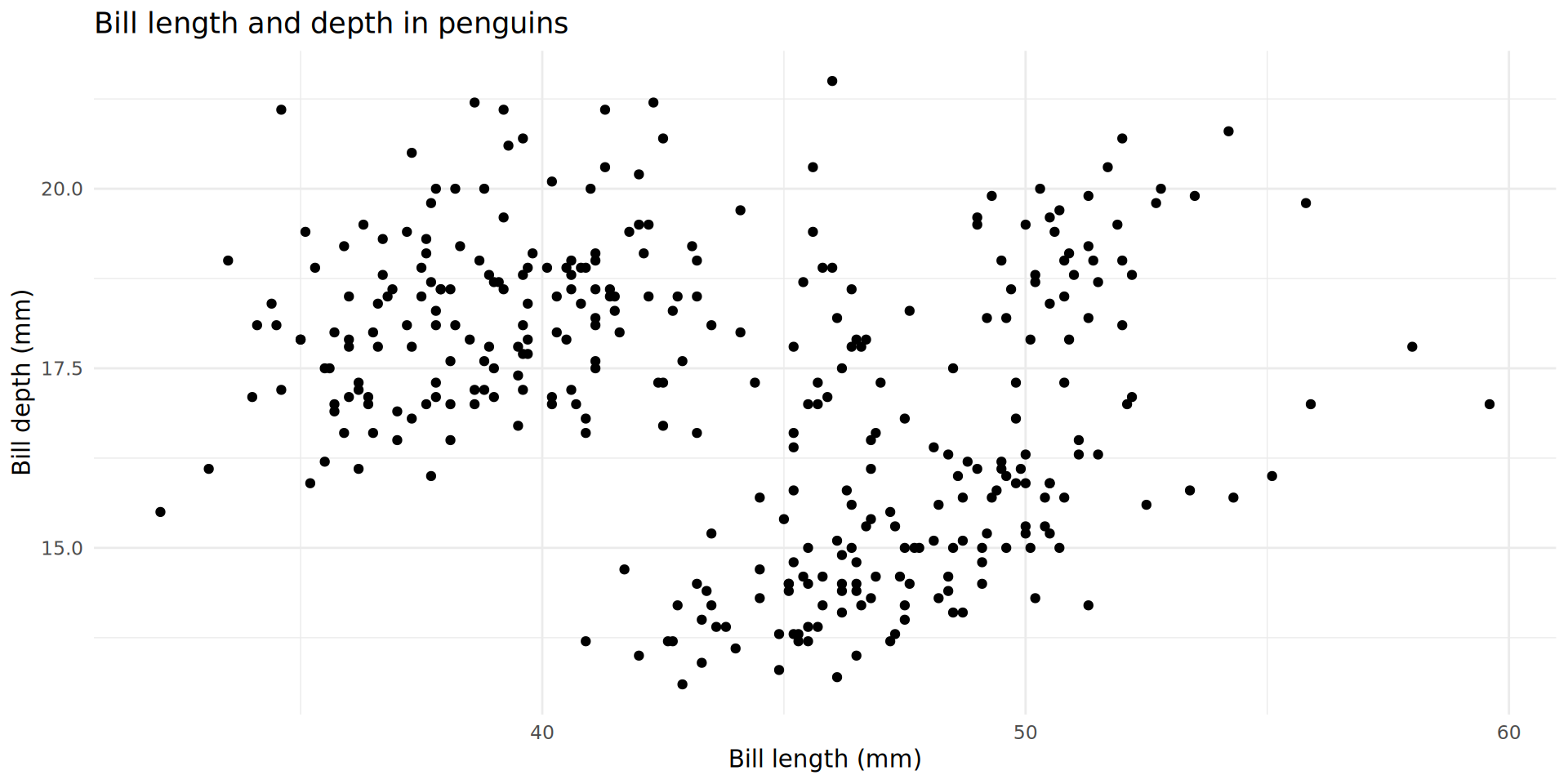
Mapping additional variables
mapping = aes() is used to map a variable to an axis.
Variables can also be mapped to axes other than x and y:
Colour, size, and shape are all axes we can map aesthetics to.
Mapping additional variables
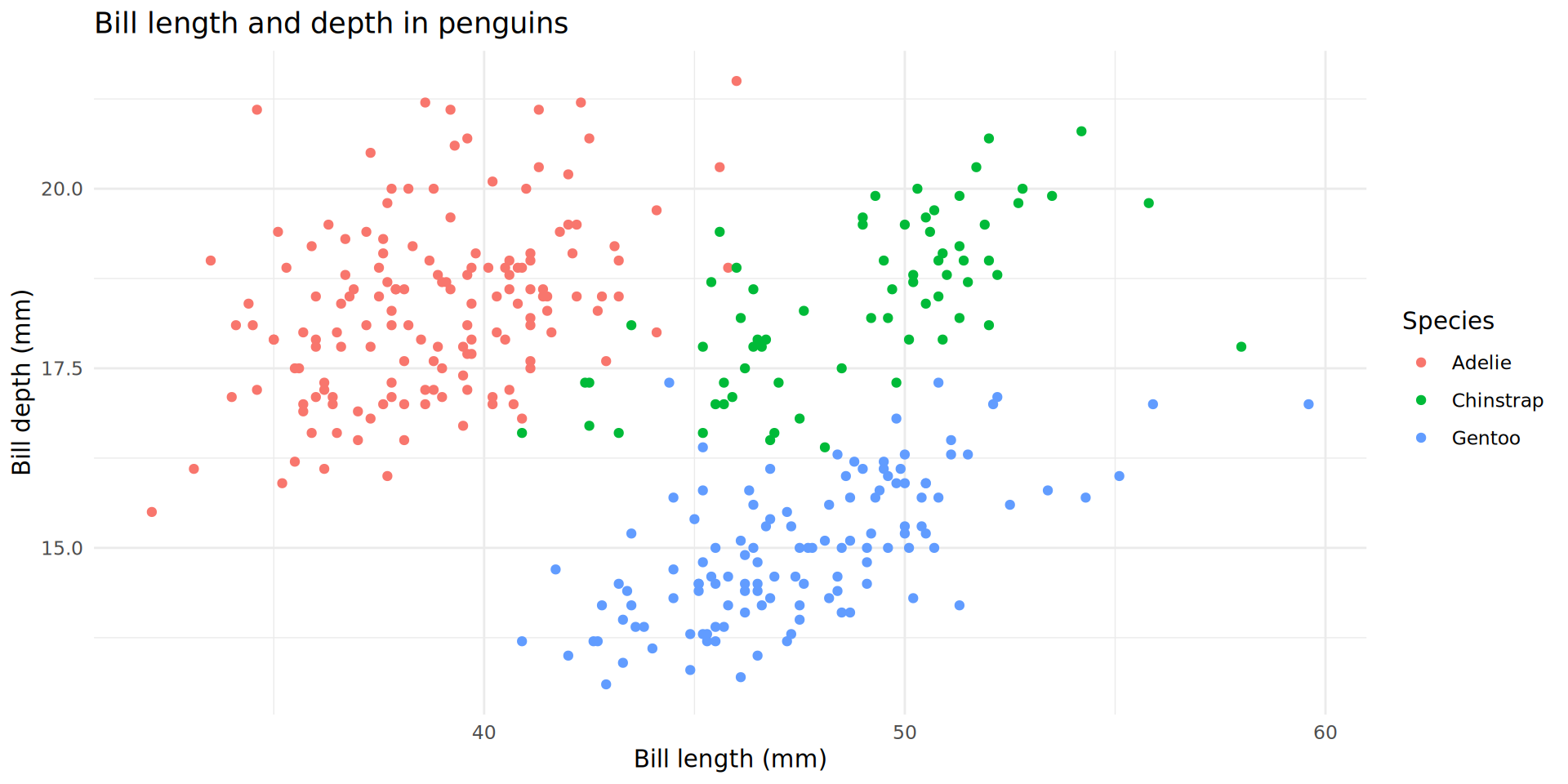
Arguments for fine control
Place arguments within the geom_ you want to control, or within theme() for overall visuals.
ggplot(data = penguins,
mapping = aes(x = bill_len,
y = bill_dep,
colour = species)) +
labs(title = "Bill length and depth in penguins",
x = "Bill length (mm)",
y = "Bill depth (mm)",
colour = "Species") +
theme_minimal() +
theme(plot.title = element_text(face = "bold")) +
geom_point(size = 3,
alpha = 0.6)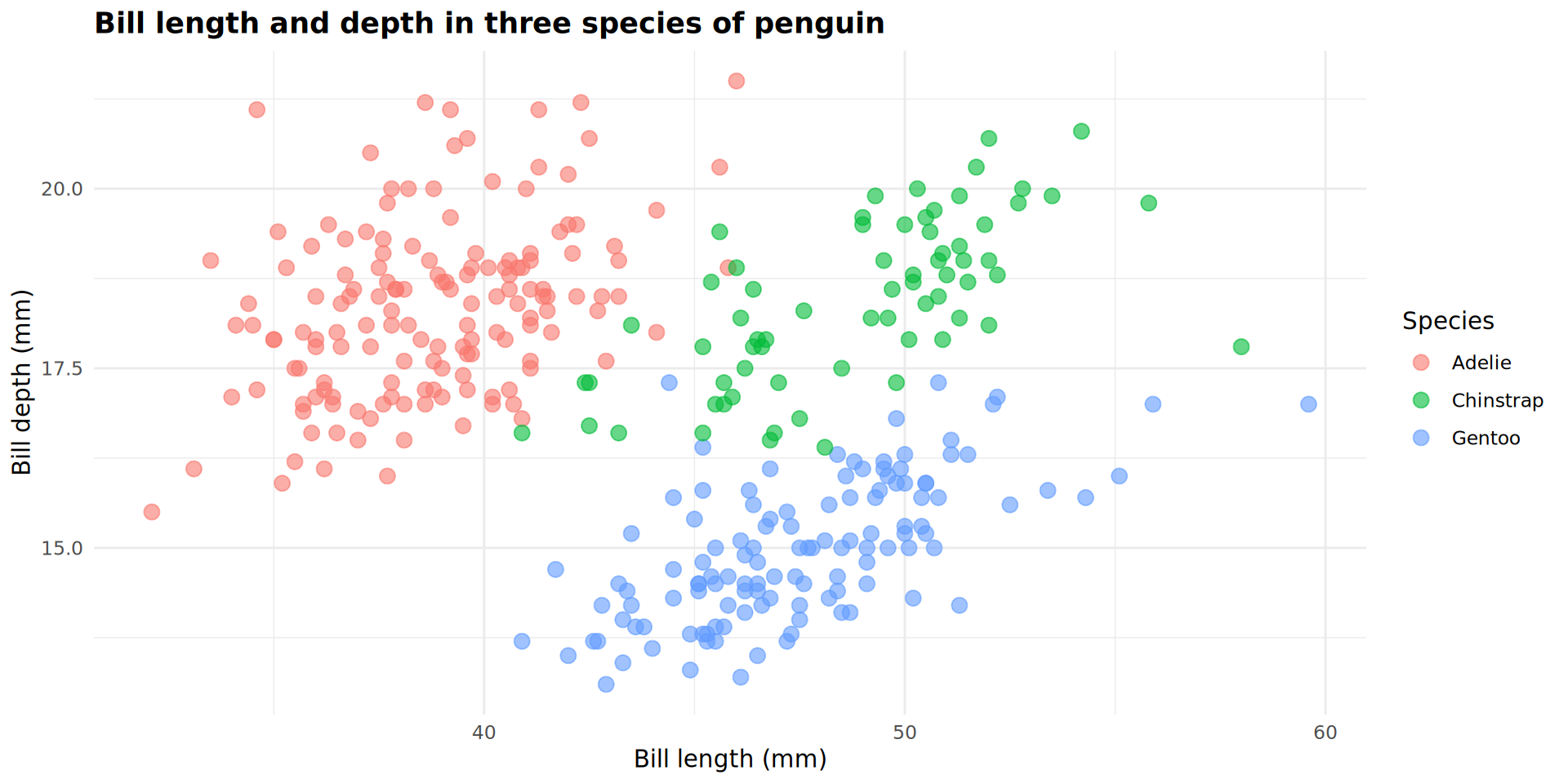
Complex arguments for geom_point()
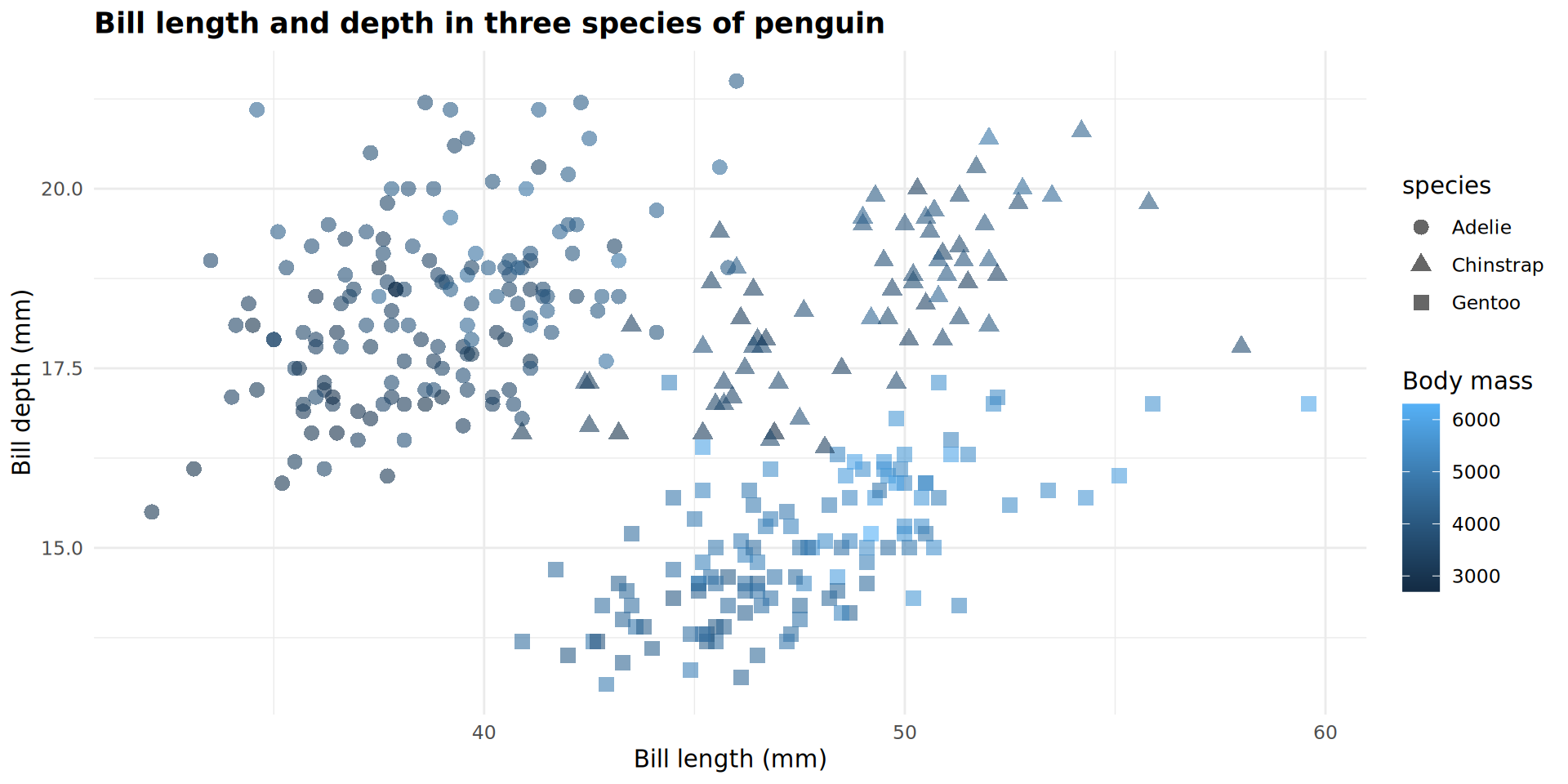
It’s your turn!
Exercise 3
How was this plot created? Use code from the previous slides and modify it yourself to duplicate this plot.
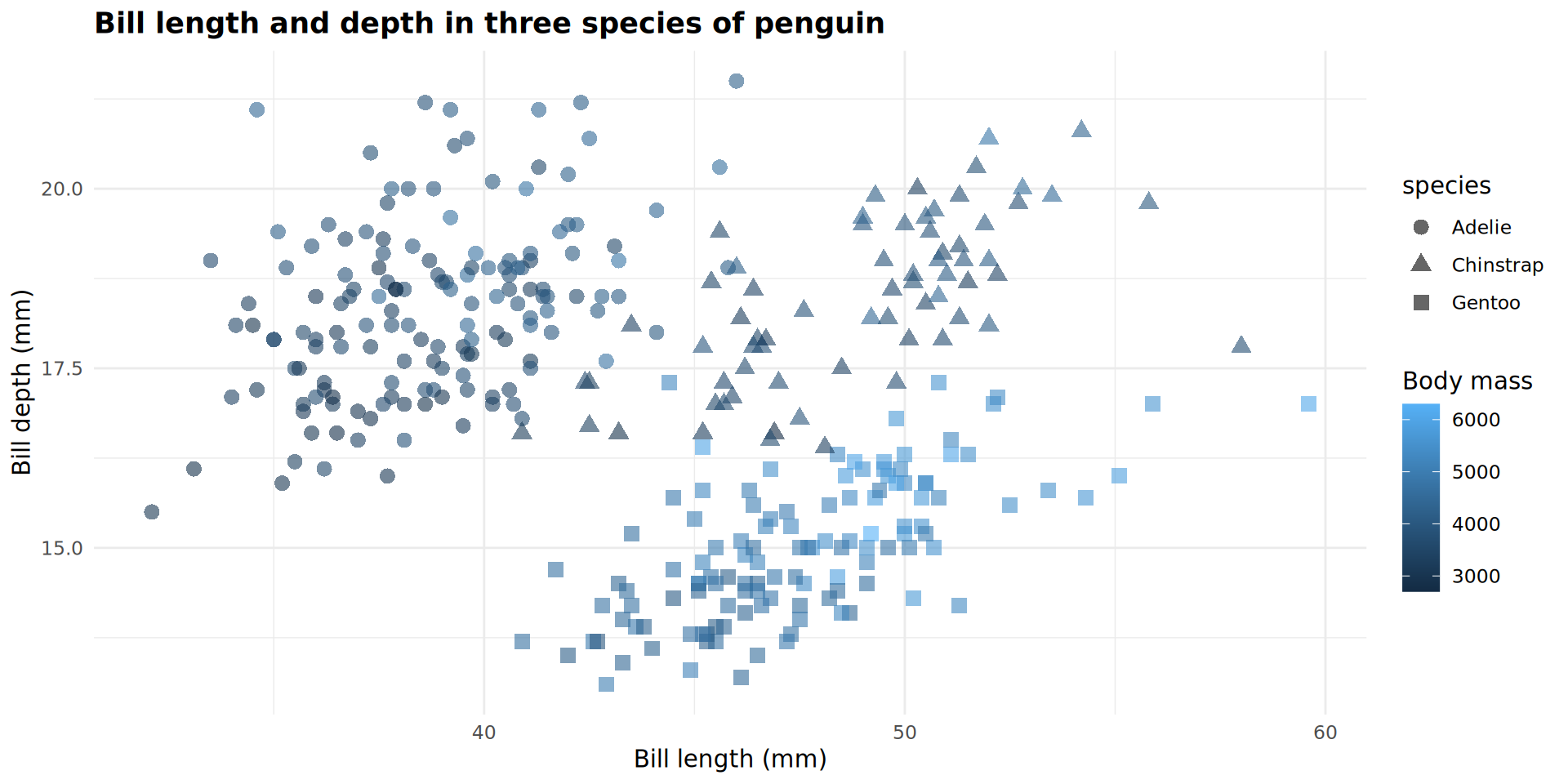
Solution and explanation
ggplot(data = penguins,
mapping = aes(x = bill_len,
y = bill_dep,
colour = body_mass,
shape = species)) +
labs(title = "Bill length and depth in three species of penguin",
x = "Bill length (mm)",
y = "Bill depth (mm)",
colour = "Body mass") +
theme_minimal() +
theme(plot.title = element_text(face = "bold")) +
geom_point(size = 3,
alpha = 0.6)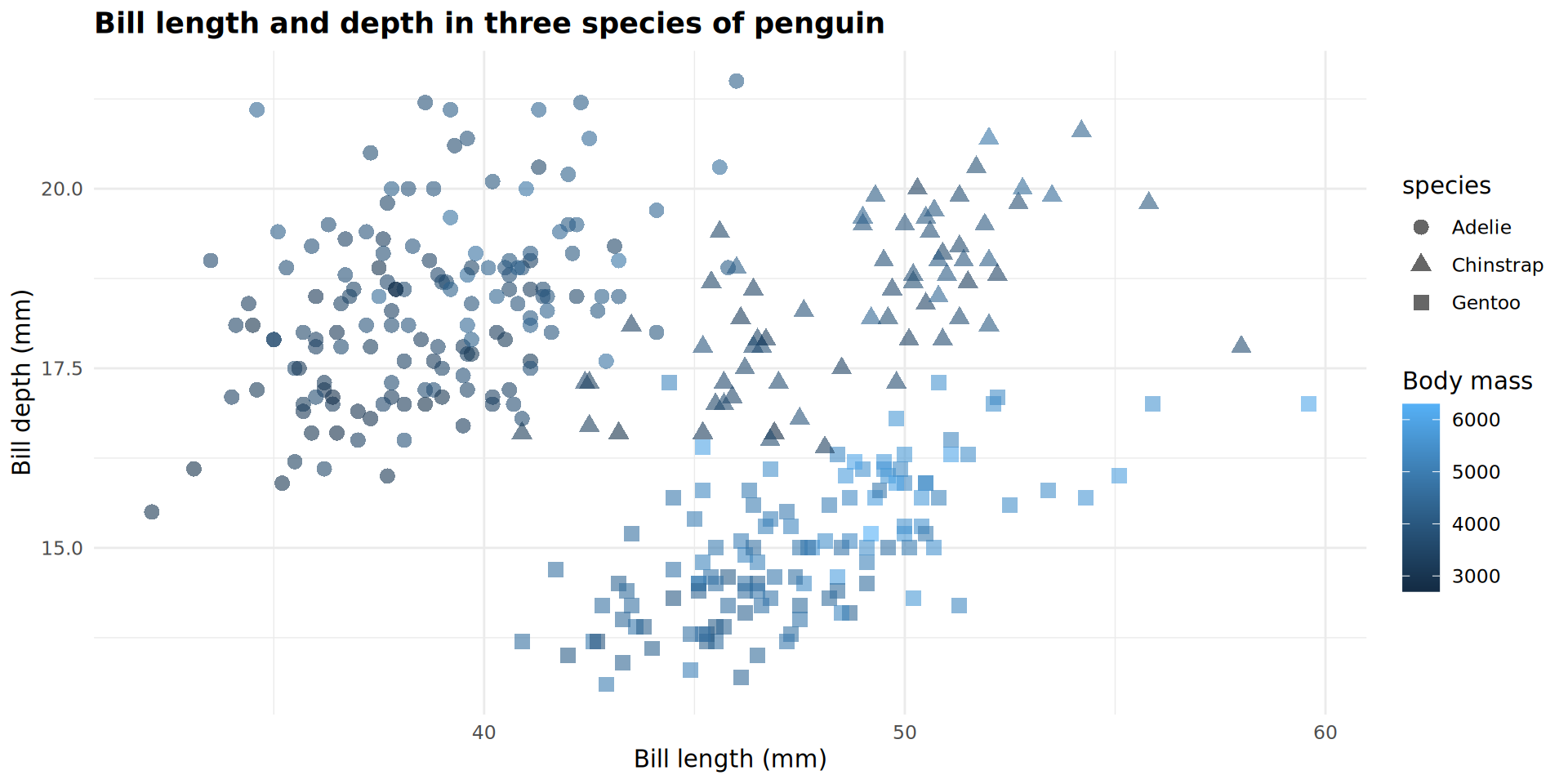
Writing in a shorter format
Because “data =” and “mapping =” are fundamental arguments, these can be shortened:
Is the same as:
Storing plot information
We can save plot information as an object.
Store mapping, theme, and other options.
Combine plot object with different geoms.
Plot objects
Plot objects
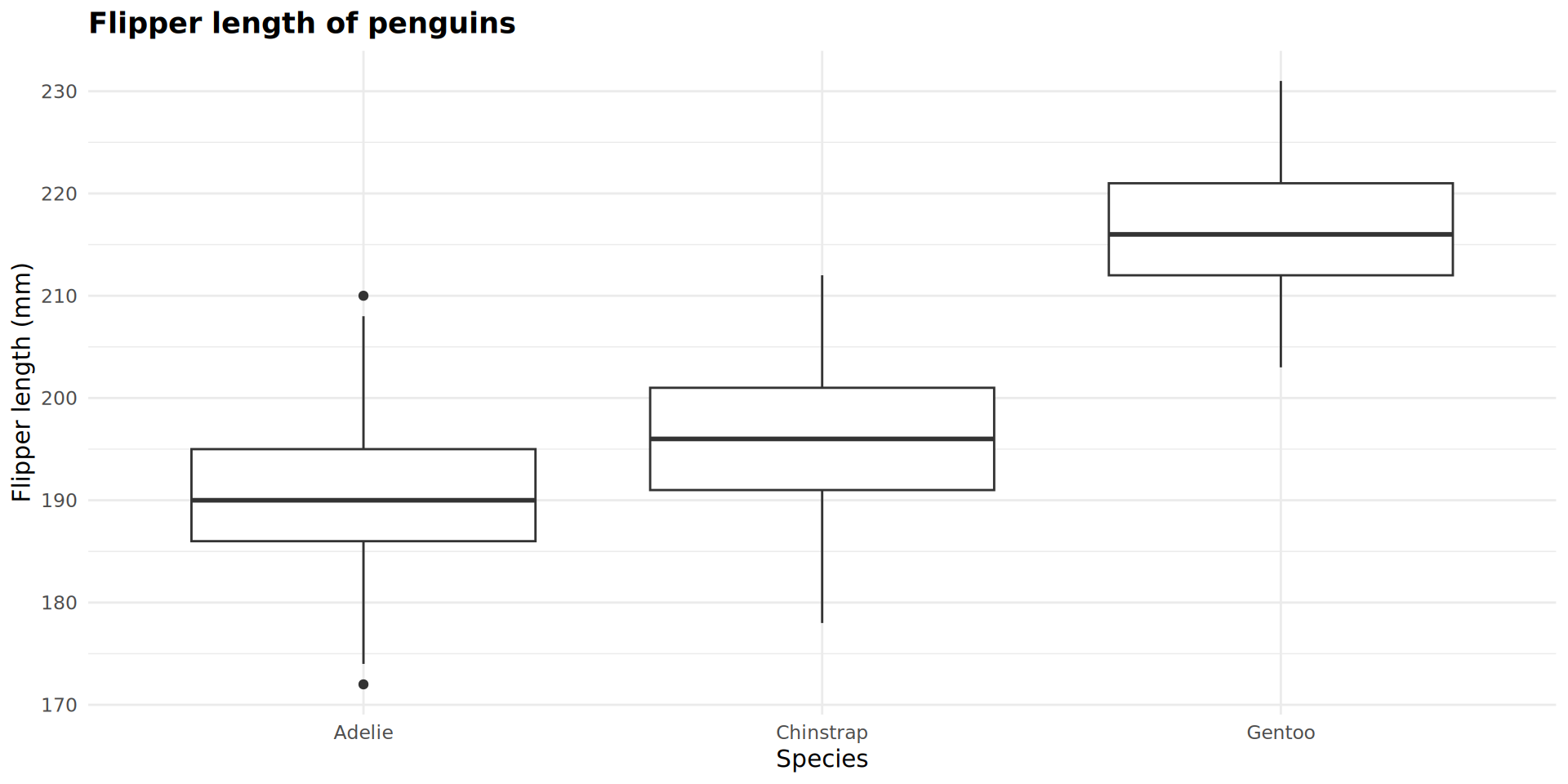
Plot objects
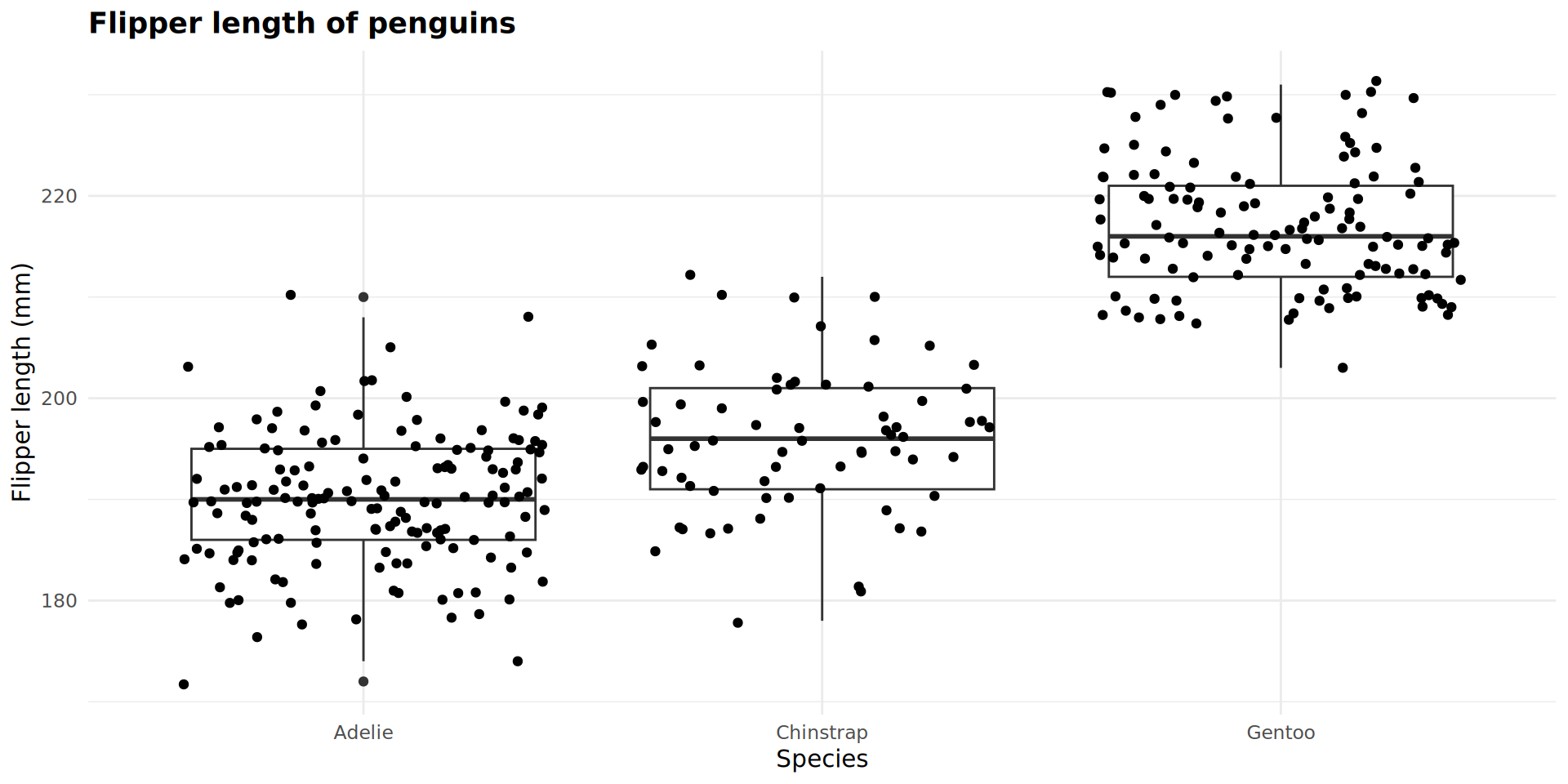
Combining geoms
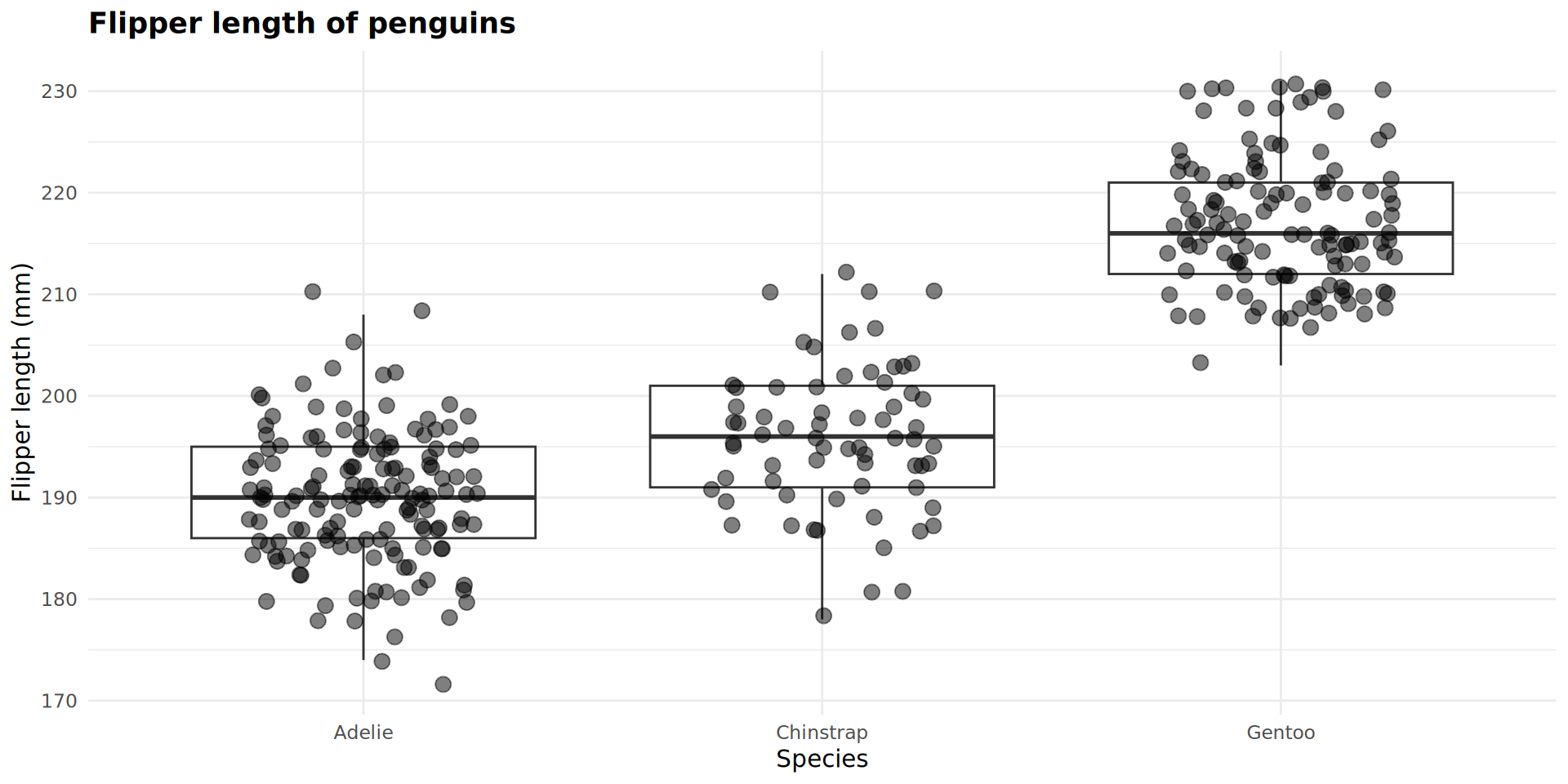
Adding variables as arguments from here
I can map variables such as species or island to the data points in geom_jitter.
Because these were not mapped in p_flipper_len, I have to use mapping = aes() to connect a variable to an aesthetic.
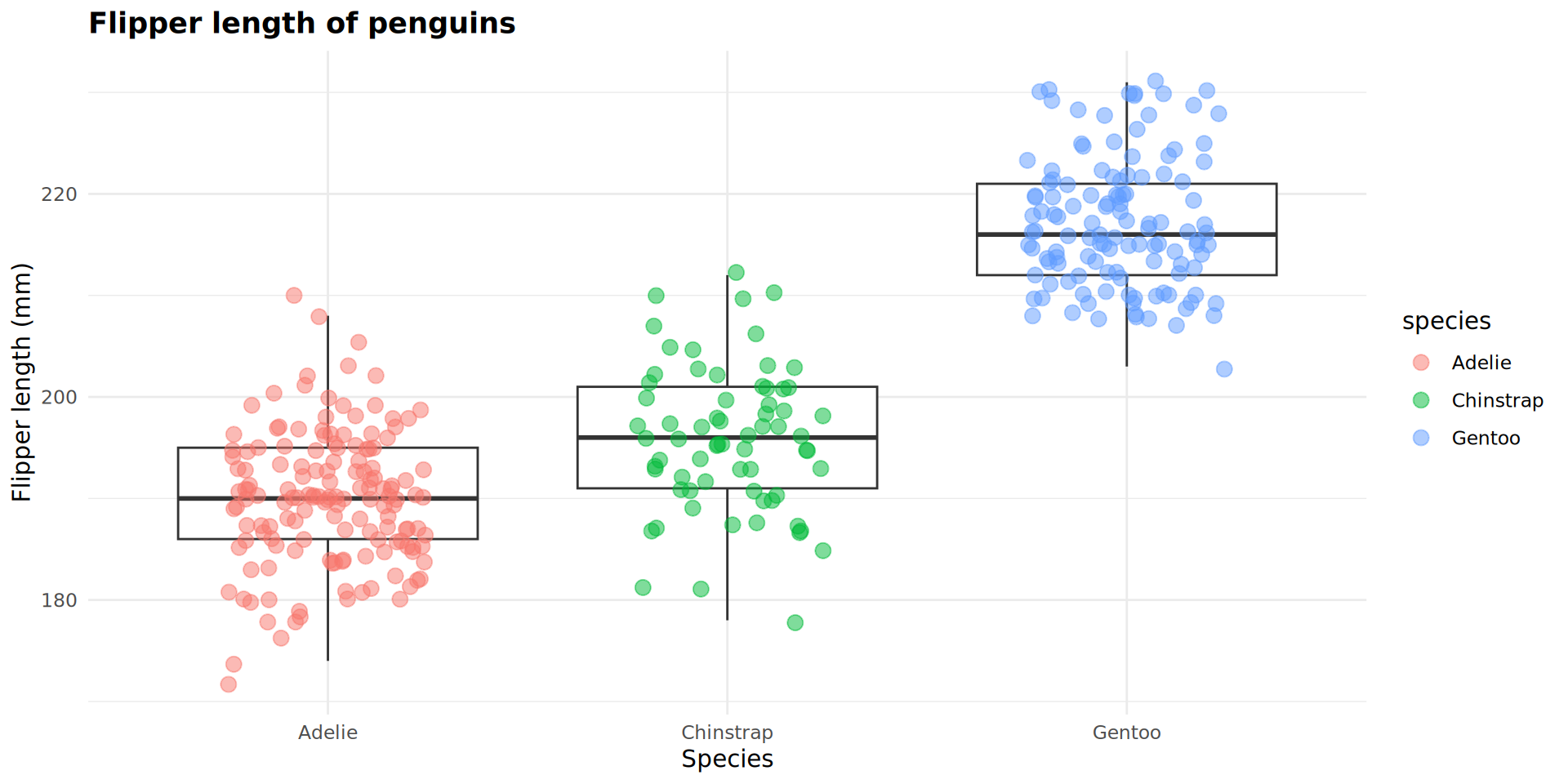
Use aes() when specifying data that is coming from an object.
Short break
Recap
How do we import data?
read_csv(),read_delim()
What functions can we use to initially look at our data?
head(),tail(),glimpse(),class(),str(), andsummary()
What are the three parts of a ggplot codeblock?
ggplot(), mapping =aes(), andgeom_X()
How do we control titles, labels, and legends?
labs()to add text,theme()to control text features
Facets and combining plots
Using facet_wrap() and facet_grid() we can split data across multiple plots.
Patterns can become clearer when data is split.
We can combine discrete plots together into a figure with the patchwork and cowplot libraries.
Join data from multiple sources to tell a cohesive story.
facet_wrap() and facet_grid()
facet_wrap() splits data according to a single variable (e.g., split samples by sex).
facet_grid() splits data according to two variables (e.g., split samples by age group and treatment group).
Preparing the data
Example with facet_wrap()
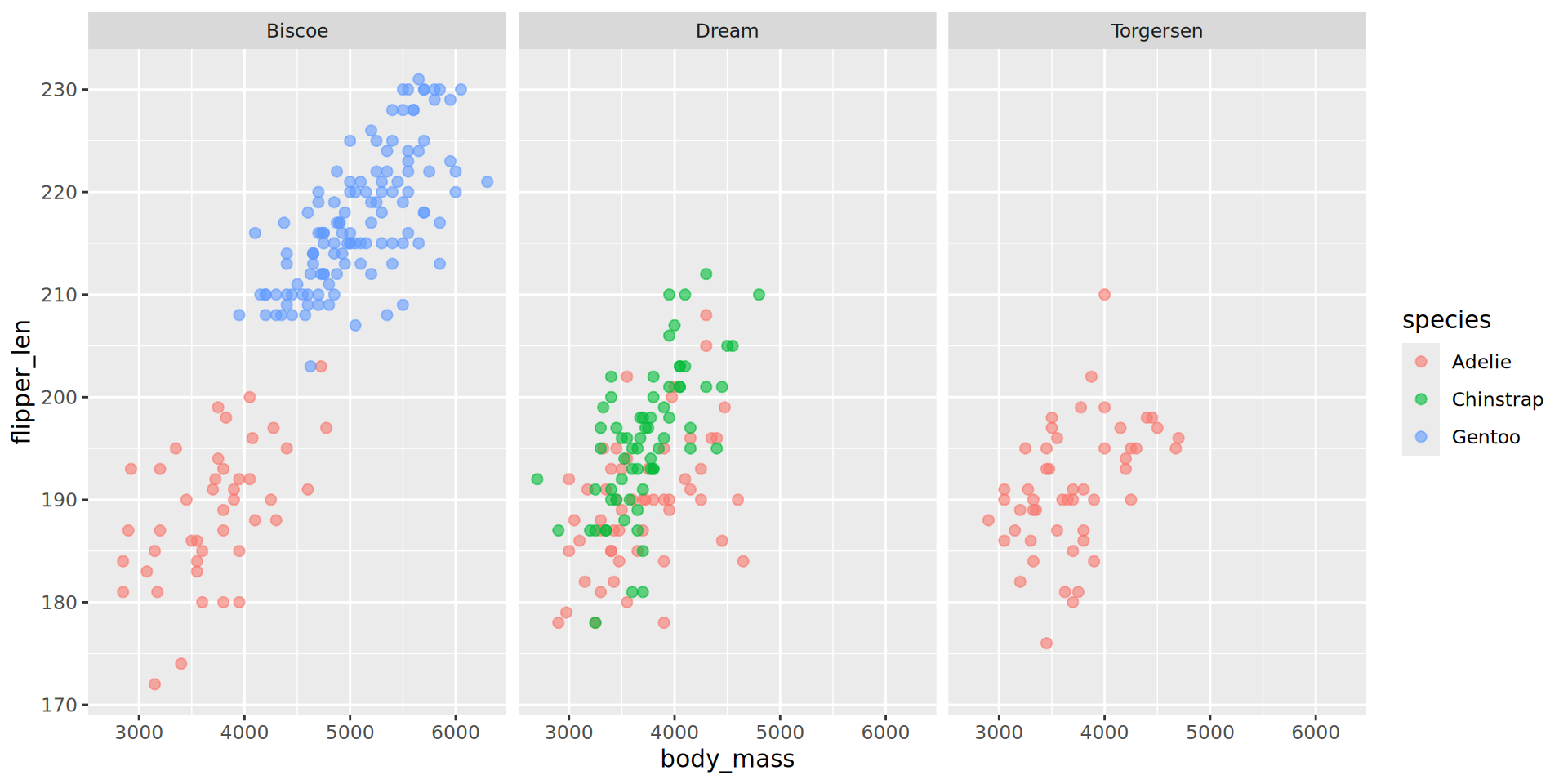
Example with facet_grid()
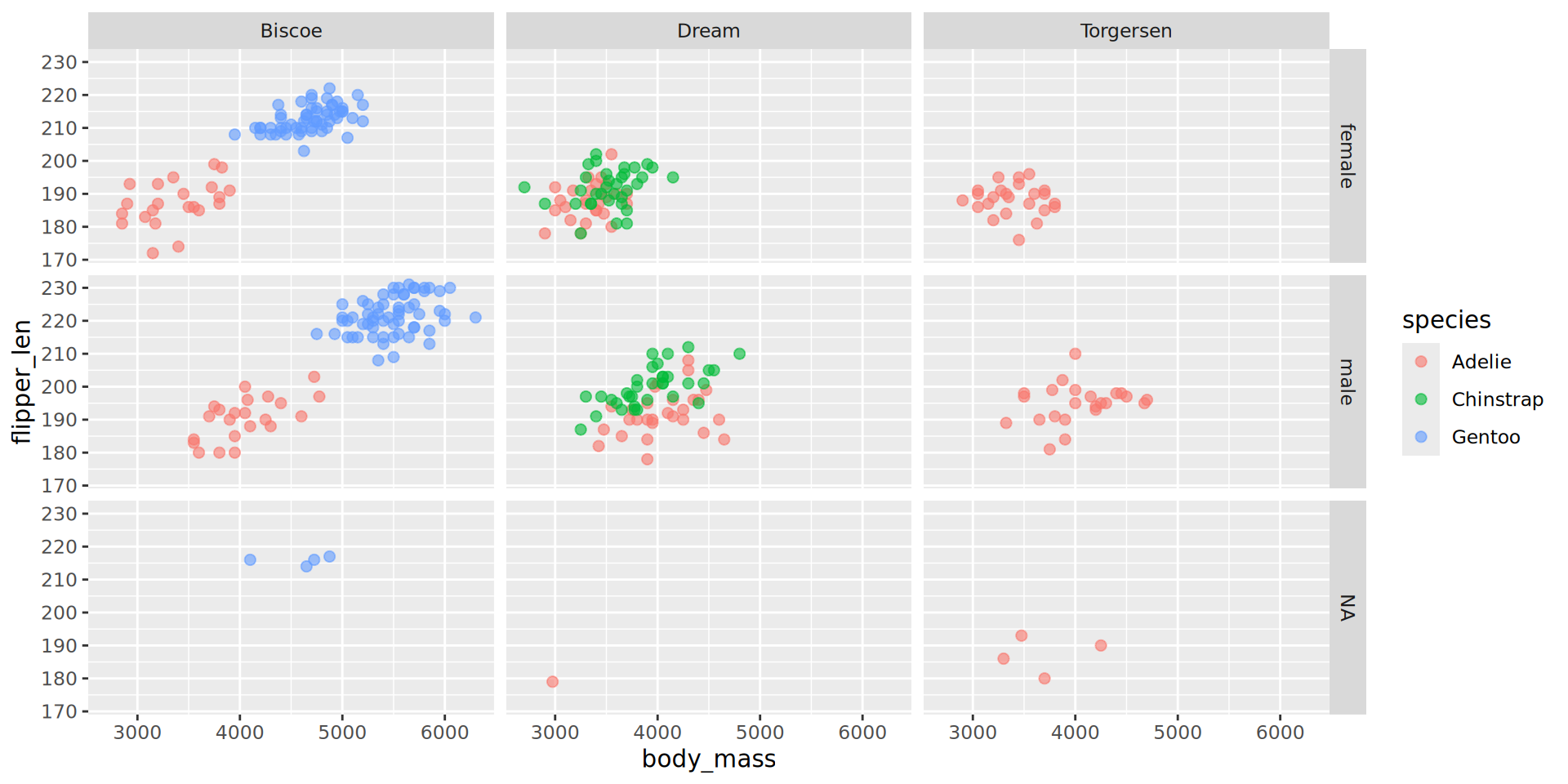
Combining plots into a single image
Two packages, patchwork and cowplot, provide the ability to combine plots.
patchwork requires stored plots
Make two plots.
p1 <- ggplot(penguins,
aes(x = bill_dep,
y = bill_len,
colour = species)) +
geom_point() +
labs(title = "Flipper length and depth",
x = "Bill depth (mm)",
y = "Bill length (mm)")
p2 <- p_bm_fl +
labs(title = "Body mass and flipper length",
x = "Flipper length (mm)",
y = "Body mass (g)") +
geom_point() Combining plots with patchwork
Combine p1 and p2 with ‘arithmetic’ operators:
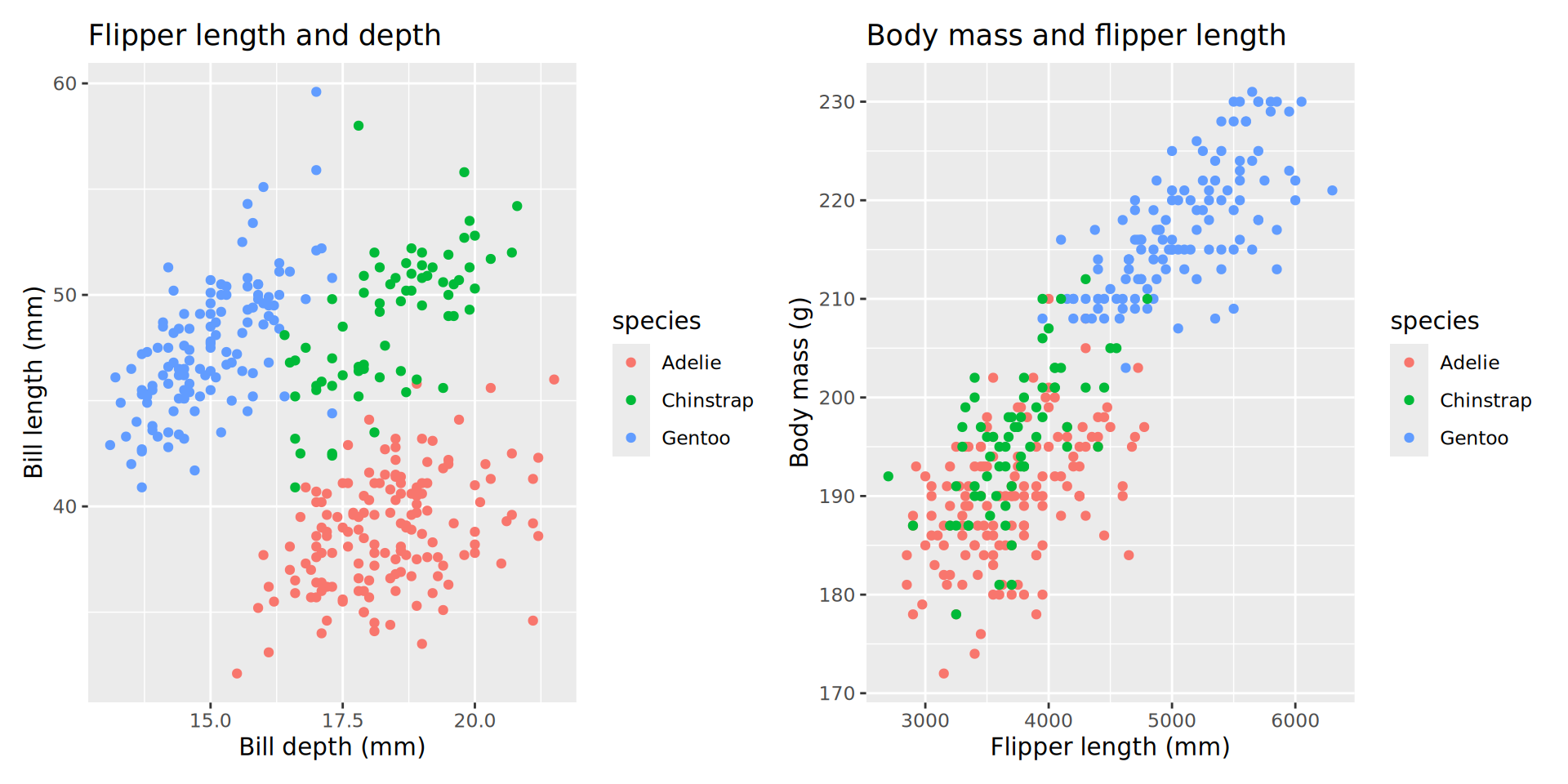
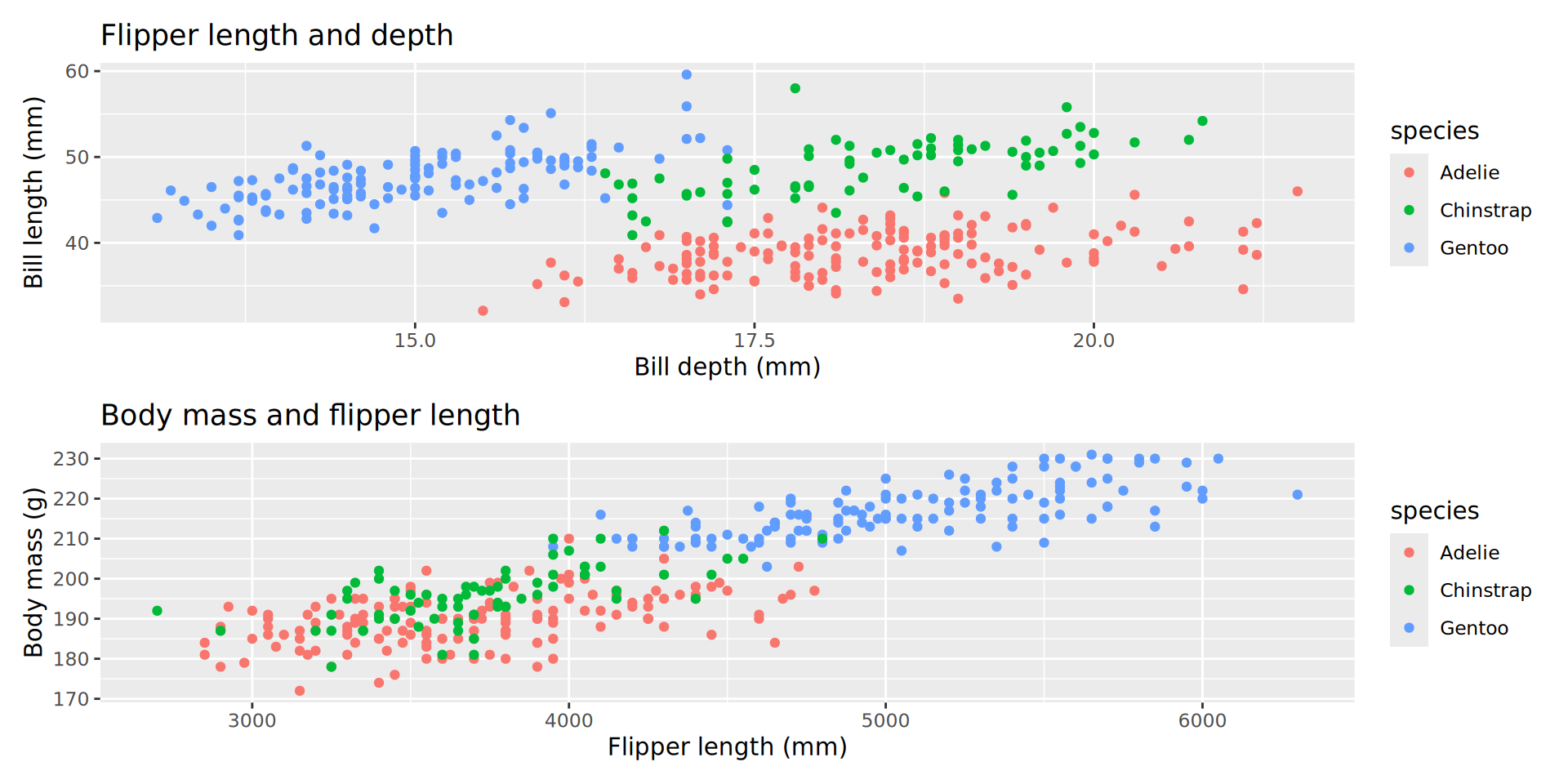
Adding trendlines (an aside)
Adding trendlines
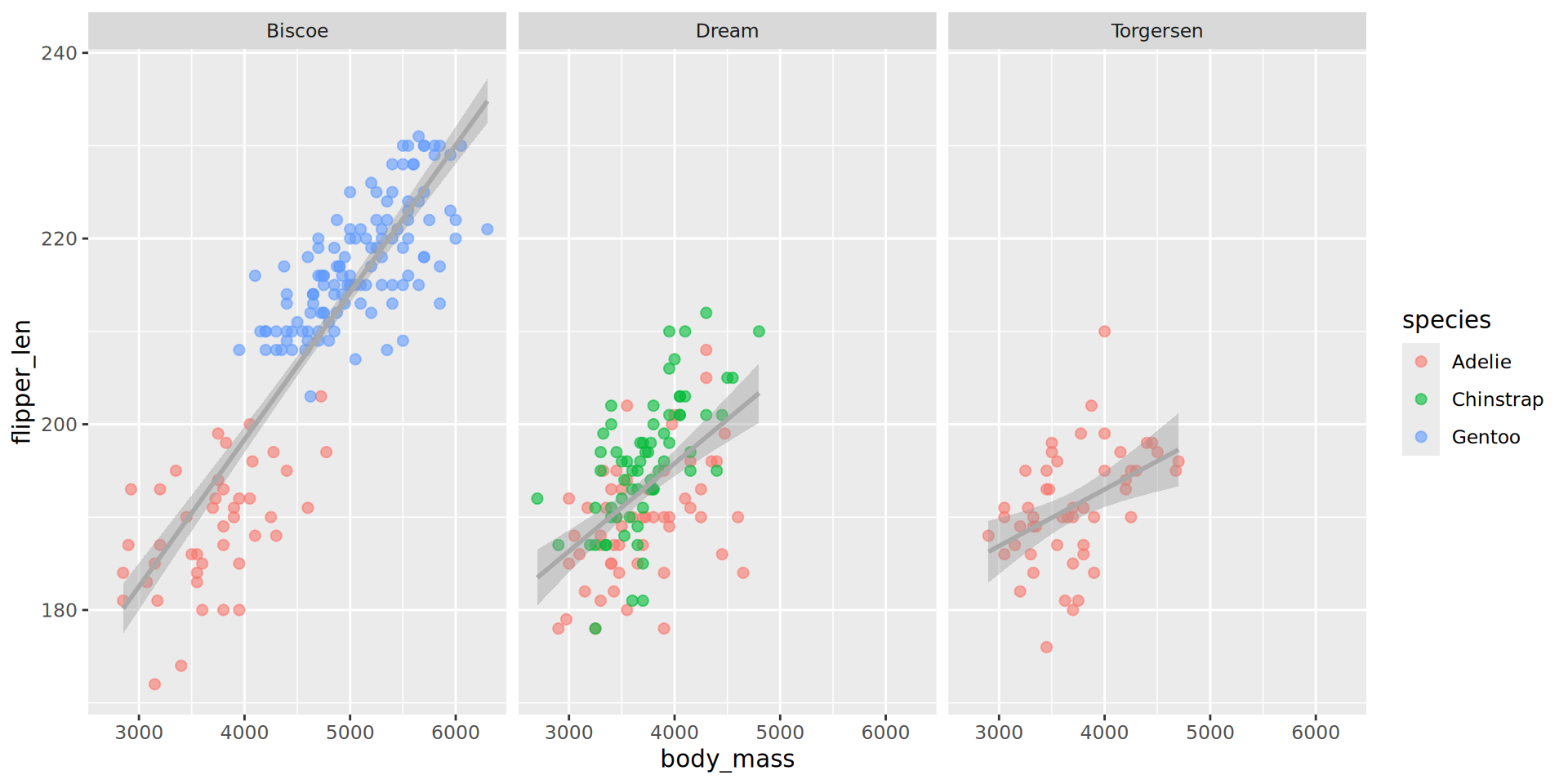
Visualising correlation with geom_tile()
Heatmaps are a useful way to visualise values across two dimensions using colour. Examples from biology are expression values for a given set of genes across samples.
This is often combined with clustering (e.g., unsupervised clustering with k-means) to identify patterns which differ between groups.
Here we will create a simple supervised tile heatmap.
Generating some example data
Here we will generate some example data showing some value (counts, words written, tasks completed etc.,) per day for five weeks.
set.seed(0982)
month_data <- tibble(
date = seq(as.Date("2024-12-01"),
as.Date("2024-12-31"),
by = "day"),
count = sample(1:20, 31, replace = TRUE) # Random counts per day
) |>
mutate(
wday = wday(date, label = TRUE, abbr = TRUE), # Day of the week (Sun-Sat)
week = (day(date) - 1) %/% 7 + 1 # Week number (1 to 5)
)
month_data |> head() # A tibble: 6 × 4
date count wday week
<date> <int> <ord> <dbl>
1 2024-12-01 2 Sun 1
2 2024-12-02 3 Mon 1
3 2024-12-03 18 Tue 1
4 2024-12-04 19 Wed 1
5 2024-12-05 8 Thu 1
6 2024-12-06 2 Fri 1Starting with geom_tile()
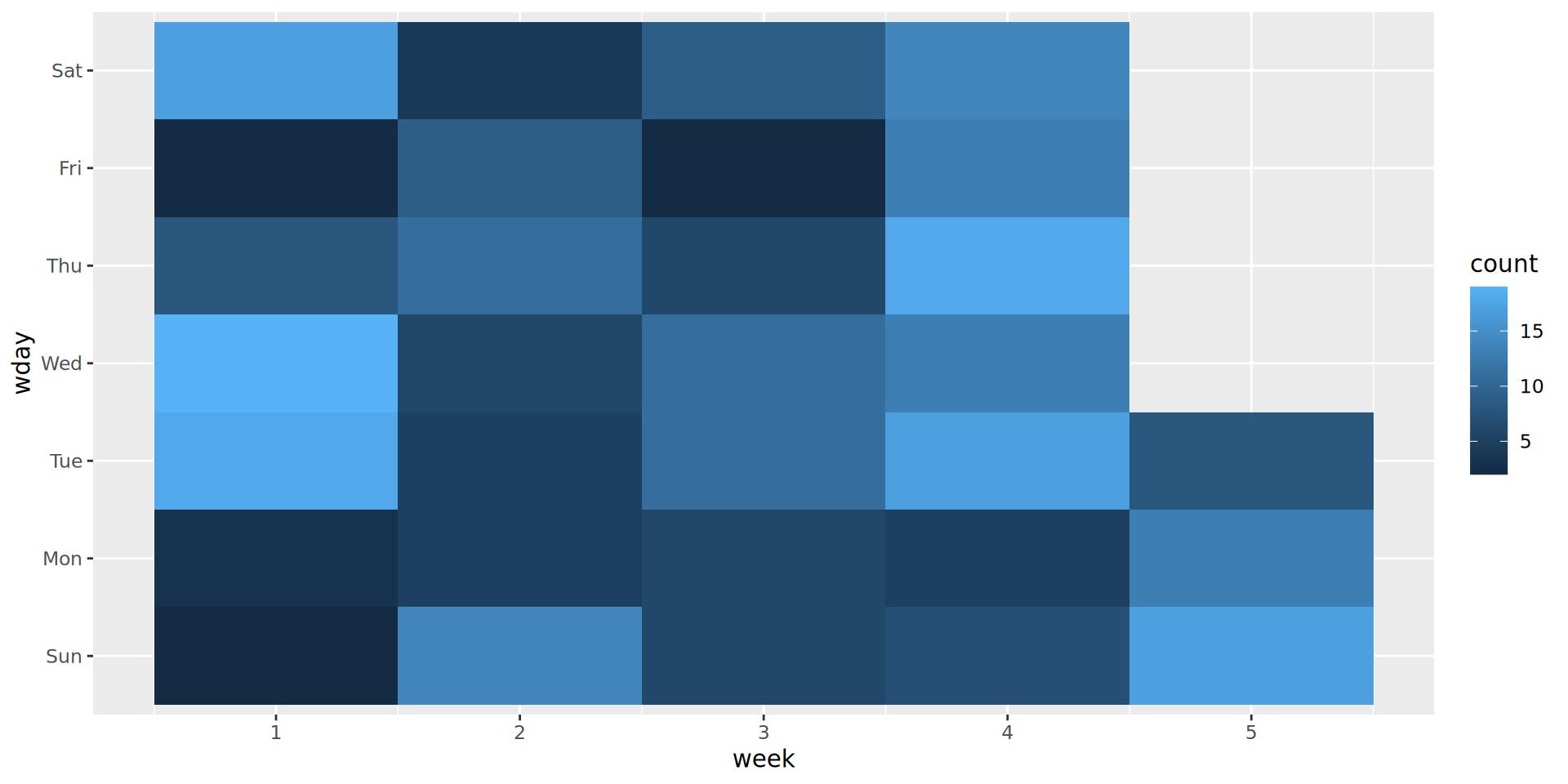
What are the issues with this plot?
Improving geom_tile()
We need to:
Change the x and y axis (putting days the y).
Invert the order of days (use the
scale_y_reverse()function).Use
labs()to add a title, axis labels and rename the fill (legend) variable.Additionally, adding colour = “white” as an argument for
geom_tile()will add separation between grid points.
Solution
Solution
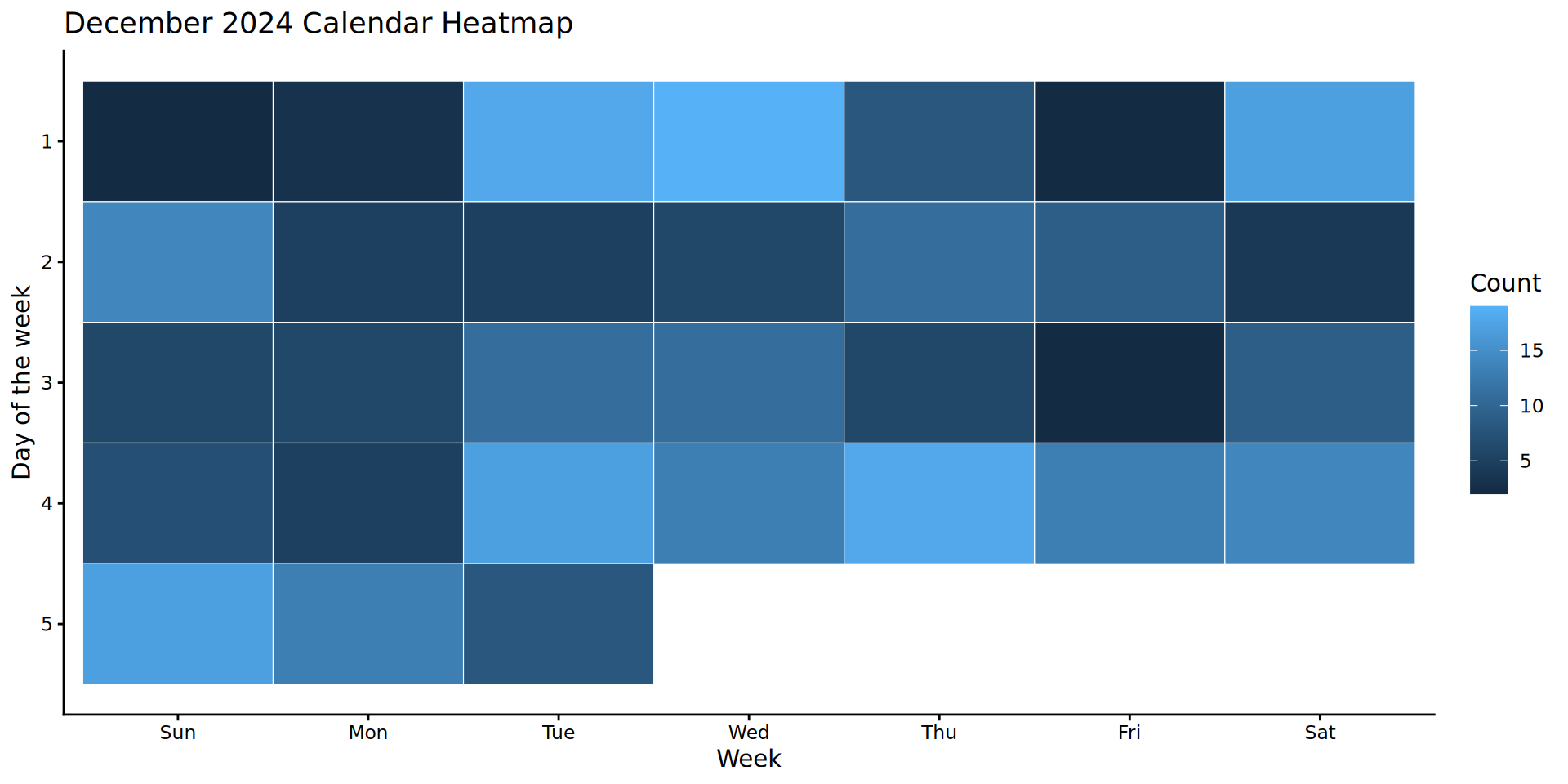
Visualising change over time (line plots)
set.seed(02) # Please set this seed for this example
# Create stock price data for three different stocks
dates <- seq.Date(from = as.Date("2024-01-01"),
by = "week",
length.out = 200)
stocks <- tibble(
date = rep(dates, times = 3),
stock = rep(c("Stock A", "Stock B", "Stock C"), each = length(dates)),
price = c(
100 + cumsum(rnorm(length(dates), mean = 0, sd = 5)), # Stock A fluctuates around 100
80 + cumsum(rnorm(length(dates), mean = 0, sd = 4)), # Stock B fluctuates around 80
120 + cumsum(rnorm(length(dates), mean = 0, sd = 6)) # Stock C fluctuates around 120
)
)Line chart with geom_line()
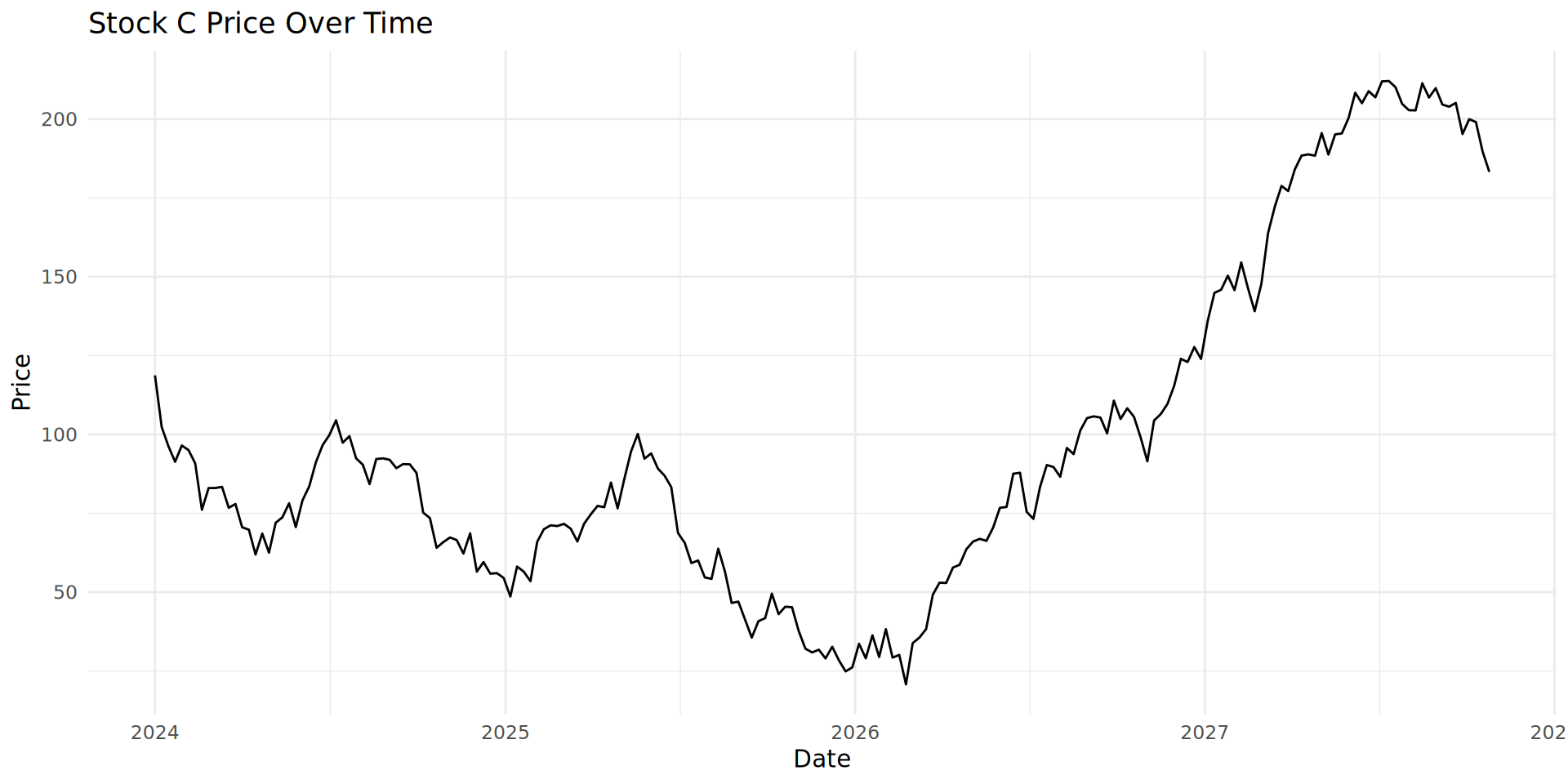
Combining geom_line() and geom_fill()
geom_area() adds a filled or shaded area. The plot looks much less empty.
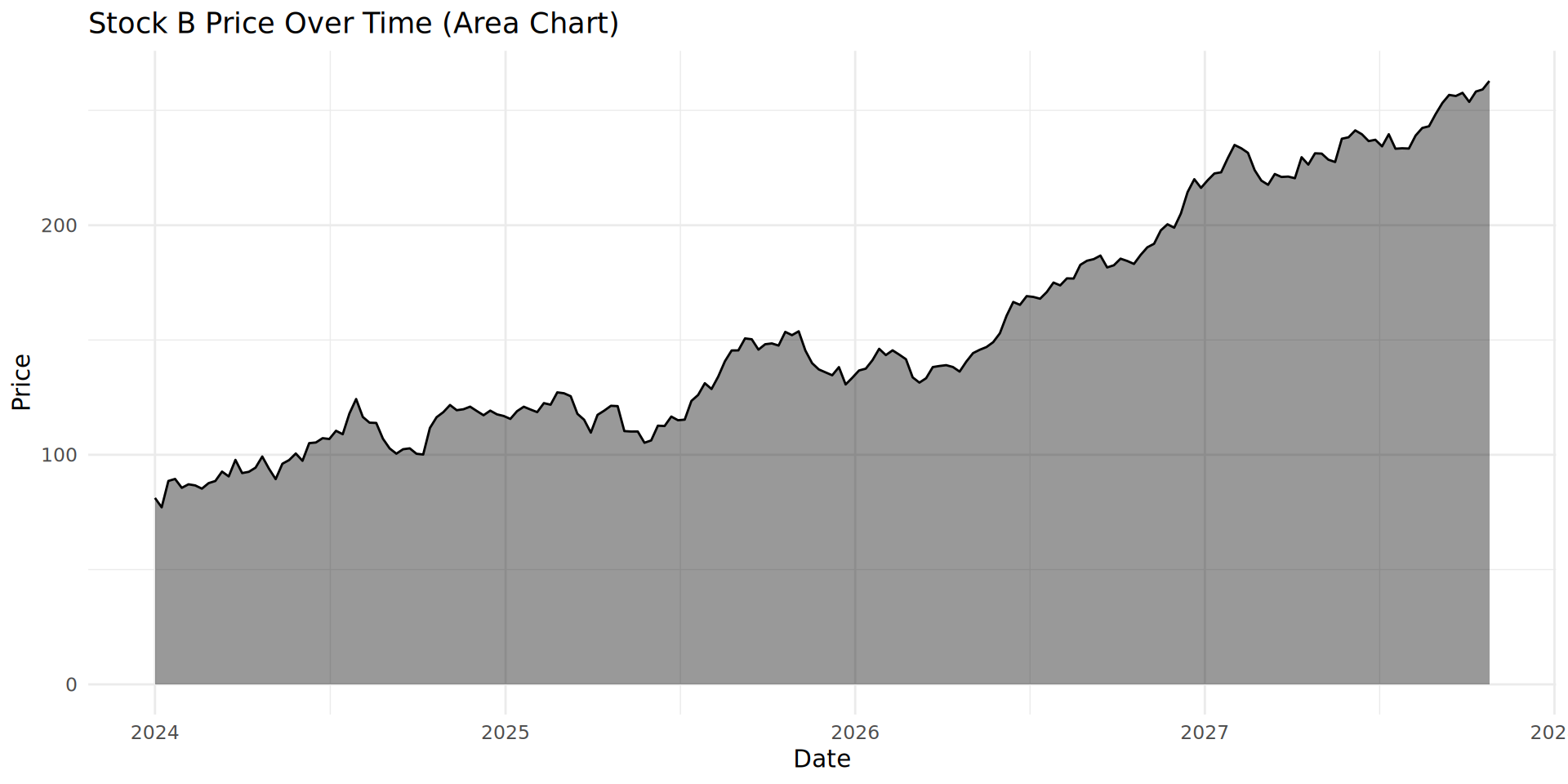
A popular option: the stacked area chart
Look at this example from an excellent data visualiser Cédric Scherer::
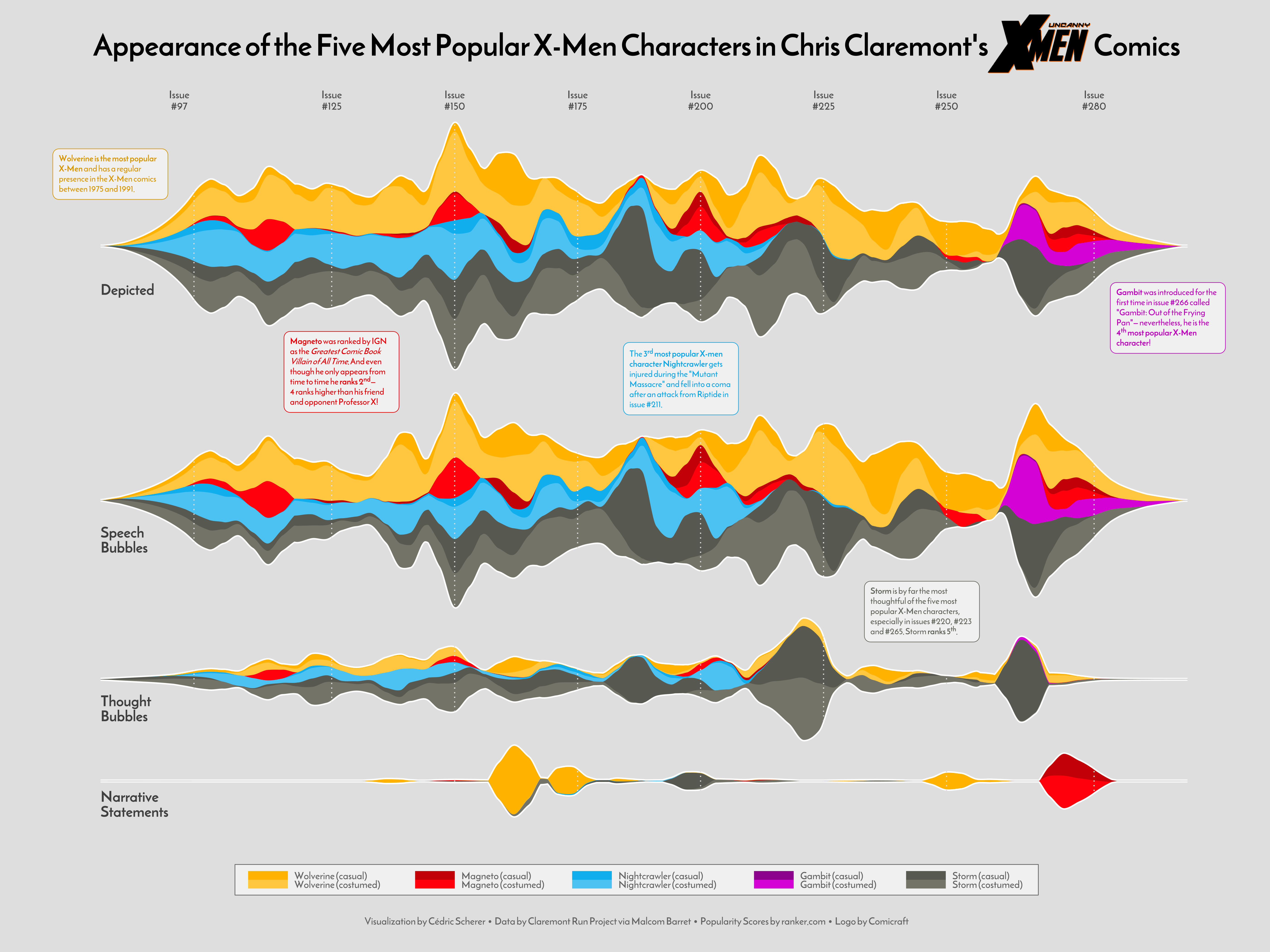
Creating a stacked area chart
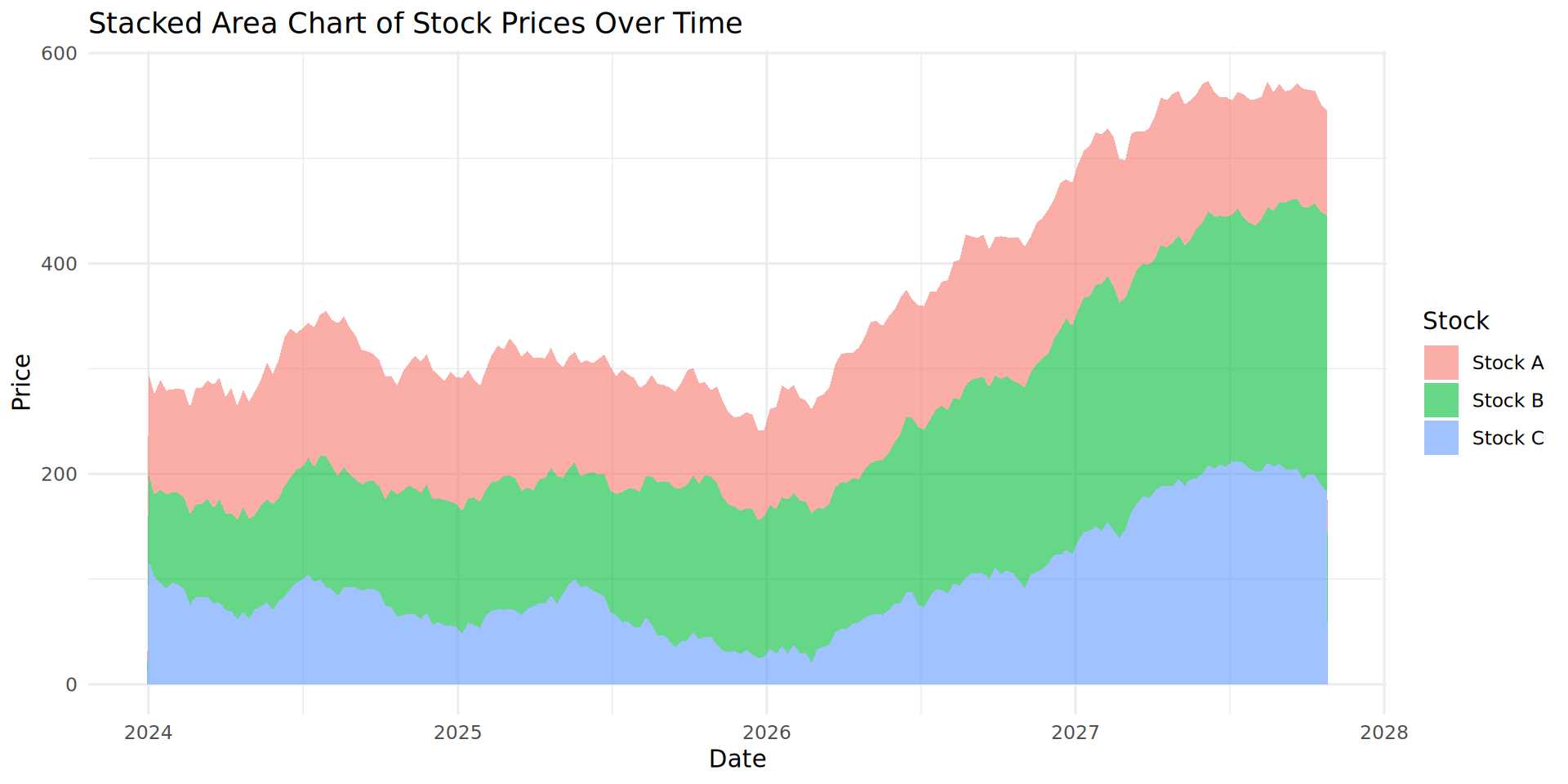
Critically analysing our stocks
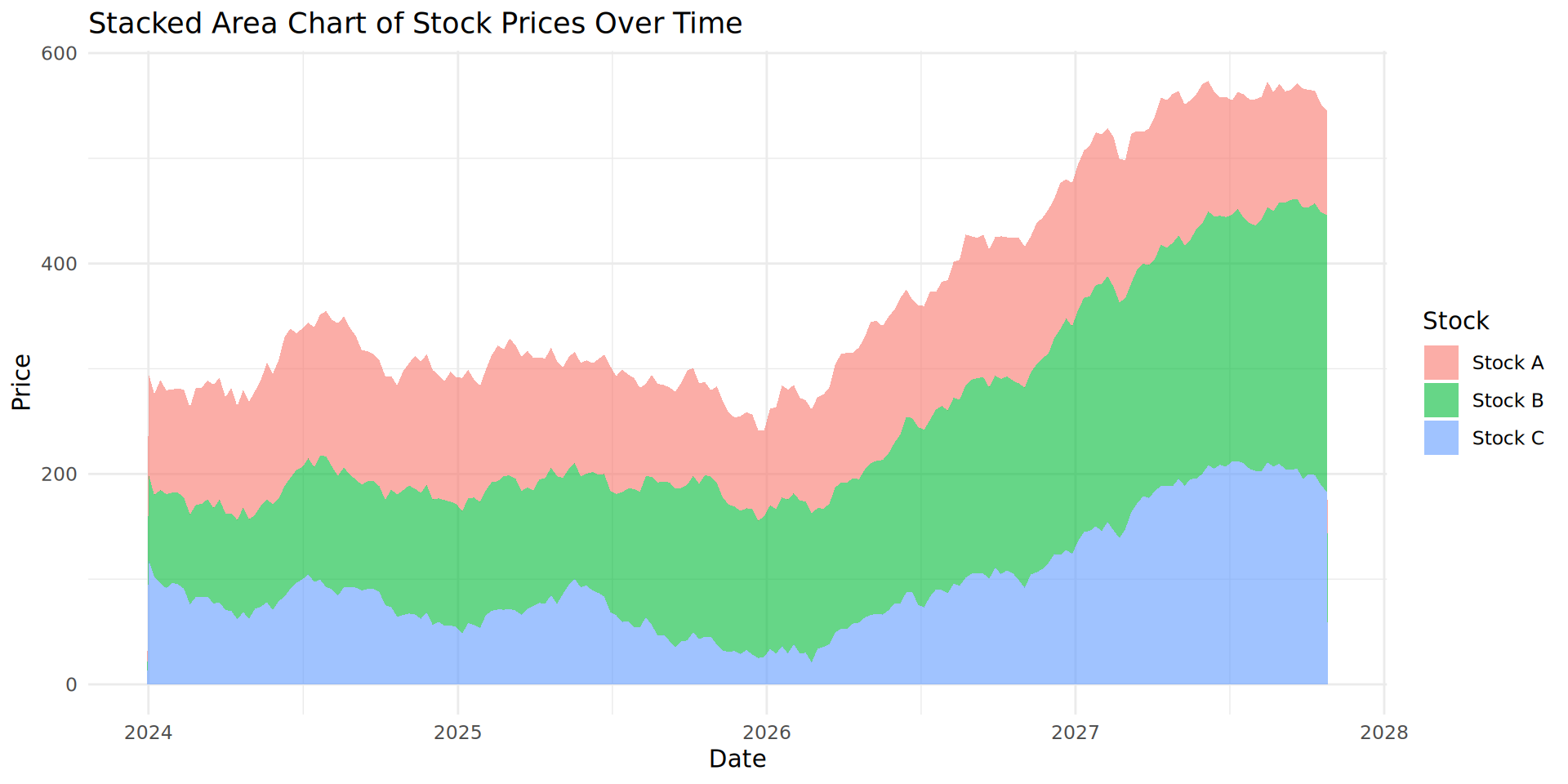
What is happening with our stock values?
Stock B and C are increasing over time. However, Stock A is not changing.
It is difficult to accurately estimate Stock A’s change in value over time because of the stacked nature of the plot.
We recommend you do not use stacked area plots, despite their popularity.
Recommended break time
The remainder of this workshop is a walkthrough of a real data visualisation.
Start to finish visualisation with tidytuesday
tidytuesday is a community initiative that publishes a weekly dataset in an easy-to-access format. Each week, people create a visualisation using the shared dataset and publish it on social media.
In 2024, tidytuesday had the aim of being featured in 10+ training courses. They hit more than 30!
In this section we will walk through a real data visualisation: importing the data from an online repo, carrying out exploratory analysis, data wrangling, visualisation, more data wrangling, and a final visualisation.
Import the tidytuesday data and exploratory analysis
Exploratory analysis
Use functions we have covered in this workshop to explore the data. Note down your initial conclusions and impressions.
Specifically:
What are five things you notice about the data?
What is one question you have about the data?
What is something you might want to investigate, visualise or learn about?
Exploratory analysis initial conclusions
The data has a lot of NAs. Concentration is almost 80% NA, Rating_Count is almost 50% NA.
There is a mix of numeric and character data.
Rating_Value looks like it’s a 0-10 scale, with a mean and median around 7.3 - 7.4
The mean and median of Rating_Count vary significantly: median of 19 and a mean of 60.
The basic structure: perfume name, brand, then values and descriptions.
Questions and assumptions: I assume Rating_Value is an average, with Rating_Count describing how many Ratings contributed to this value.
Brand, Rating_Value, and Rating_Count
Aim: generate a visualisation which will show which brands consistently have high ratings for their different products, with the intention that a person with little to no knowledge about perfume (like myself) can get an idea of reliable brands.
Exercise: Mentally visualise what this figure might look like. What are the key components we will need to convey?
Assessing the means
# A tibble: 6 × 2
Brand avg_Rating
<chr> <dbl>
1 Natura 10
2 Sarahs Creations 10
3 mesOud 9.47
4 Bourjois 9.4
5 Jehanne Rigaud 9.3
6 Max Joacim 9.3
Does this mean Natura and Sarahs Creations are the best choice? Why?
- Does
summary()change your opinion?
Hypothesis
Hypothesis: some brands have very high average ratings but this is an artifact due to low sampling numbers.
i.e., Brands with more ratings are (unfairly) penalised as their average ratings are pulled downwards by individual preferences. Natura and Sarahs Creations were possibly rated by 1 - 2 people who scored the perfumes with 10s.
- How can we test this hypothesis with a visualisation?
Perfumes with fewer ratings have high variation in rating value
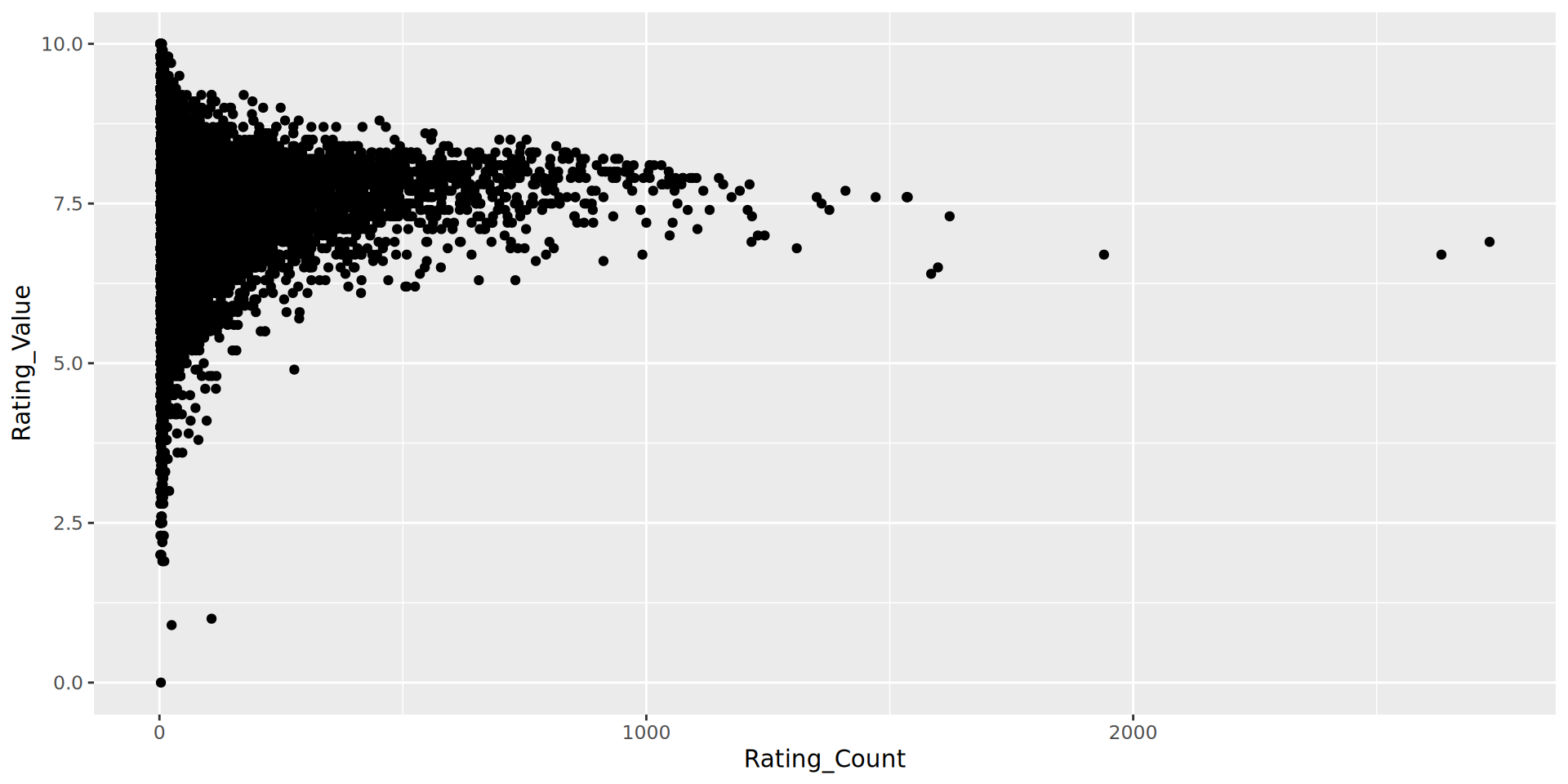
- Comment on the fact that there are no perfumes with more than 1000 ratings and a rating of less than 5. What could cause this?
An aside: being aware of other relationships
Hypothesis: There will be a significant relationship between Rating_Count and Release_Year.
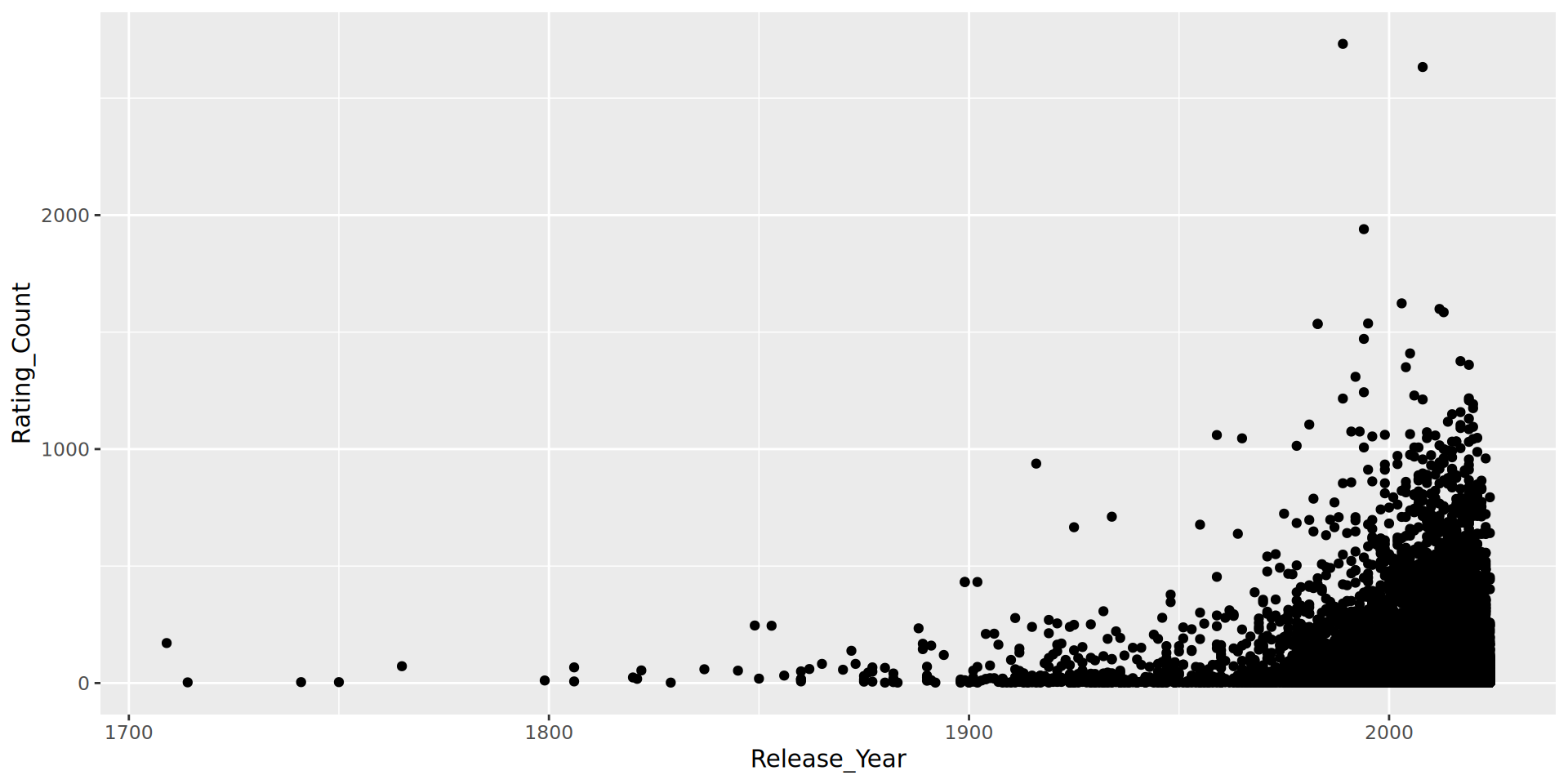
This is important because if we are filtering based on a Rating_Count threshold, we need to be aware this is reducing the likelihood of older Brands appearing.
note: I have suppressed the warnings about NAs, and will continue to do so for all future plots.
Ready to wrangle
At this point I have performed a basic exploratory analysis. I believe I understand the basics of my data and I’m ready to begin data wrangling.
For each of the data wrangling steps we will discuss the logic.
Data wrangling steps
Filtering for low rating counts: Perfumes that were rated fewer than 19 times (the median number of ratings) were removed from the data. Having a small number of ratings can skew the rating result (e.g., a single 10/10 is not representative of the perfumes quality).
Calculate the average rating by brand and the number of perfumes per brand: Perfumes were grouped by brand, and the average (mean) rating for each brand was calculated. Some brands had only a small number of perfumes, which led to skewed averages.
Data wrangling cont.
Remove brands with a small number of perfumes: Brands with fewer than 20 perfumes were removed from the analysis.
Store the perfume_brand_data object: A dataframe of 1635 rows, consisting of only perfumes that meet the above two criteria (individually rated more than 19 times, and from a brand with 20 or more perfumes).
Create and store the brand_rating_data object: A dataframe of 20 rows and three variables, consisting of the 20 brands with the highest average rating (the mean rating across all perfumes for the brand), as well as the number of perfumes in each brand.
Explore the new objects
Explore the new objects
Number Name Brand Release_Year Concentration Rating_Value
1 150 : Contemporary Clive Christian 2022 <NA> 8.1
2 150 : Timeless Clive Christian 2022 <NA> 8.1
3 1872 Acacia Clive Christian 2017 <NA> 7.9
4 1872 Basil Clive Christian 2018 <NA> 7.8
5 1872 Bergamot Clive Christian 2017 <NA> 8.1
6 1872 for Men Clive Christian 2001 <NA> 8.2
Rating_Count Main_Accords
1 58 Fruity, Fresh, Spicy, Green, Citrus
2 41 Citrus, Fresh, Spicy, Green, Woody
3 15 Floral, Fresh, Green, Woody, Spicy
4 48 Green, Spicy, Fresh, Woody, Citrus
5 14 Fresh, Spicy, Citrus, Woody, Fruity
6 407 Green, Fresh, Spicy, Citrus, Floral
Top_Notes
1 <NA>
2 <NA>
3 Black cherry
4 Spearmint, Basil, Black pepper, Nutmeg
5 Bergamot, Mandarin orange, Rosemary, Lavender
6 Galbanum, Petitgrain, Bergamot, Lime, Grapefruit, Lavender, Mandarin orange, Peach, Pineapple, Rosemary, Nutmeg, Pepper
Middle_Notes
1 <NA>
2 <NA>
3 Acacia, Freesia, Ginger, May rose
4 Clary sage
5 Clary sage, Neroli
6 Clary sage, Cyclamen, Tagetes, Freesia, Jasmine
Base_Notes Perfumers
1 <NA> Angela Stavrevska
2 <NA> Julie Pluchet
3 Amber, Cedarwood, Patchouli <NA>
4 Woods, Iris <NA>
5 Patchouli <NA>
6 Cedar, Musk, Amber, Labdanum, Patchouli, Frankincense Geza Schön
URL
1 https://www.parfumo.com/Perfumes/Clive_Christian/150-contemporary
2 https://www.parfumo.com/Perfumes/Clive_Christian/150-timeless
3 https://www.parfumo.com/Perfumes/Clive_Christian/1872_Acacia
4 https://www.parfumo.com/Perfumes/Clive_Christian/1872_Basil
5 https://www.parfumo.com/Perfumes/Clive_Christian/1872_Bergamot
6 https://www.parfumo.com/Perfumes/Clive_Christian/1872-for-men [1] "Number" "Name" "Brand" "Release_Year"
[5] "Concentration" "Rating_Value" "Rating_Count" "Main_Accords"
[9] "Top_Notes" "Middle_Notes" "Base_Notes" "Perfumers"
[13] "URL" Explore the new objects
Building a visualisation - sketching
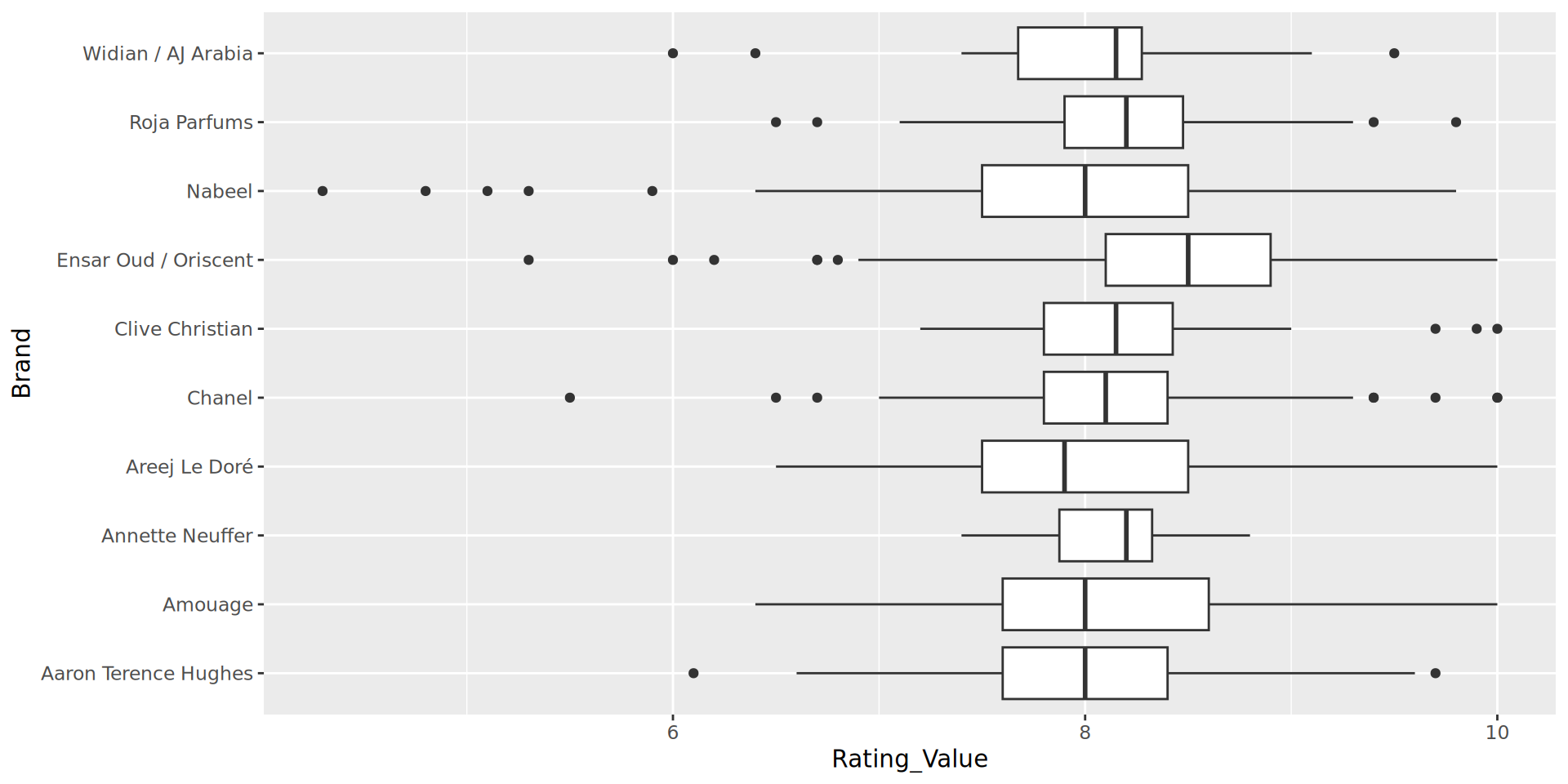
“Sketching” options
ggplot allows us to make quick plots to visualise data. For a complex task, it’s recommended to try different geoms and formats, experimenting with mappings to identify the clearest way to represent the data.
In this example I eventually discarded the boxplot. I could not see an easy way to represent the number of perfumes per brand, nor the number of ratings per perfume, and I felt this was important data.
geom_point and the brand rating average
geom_point and the brand rating average
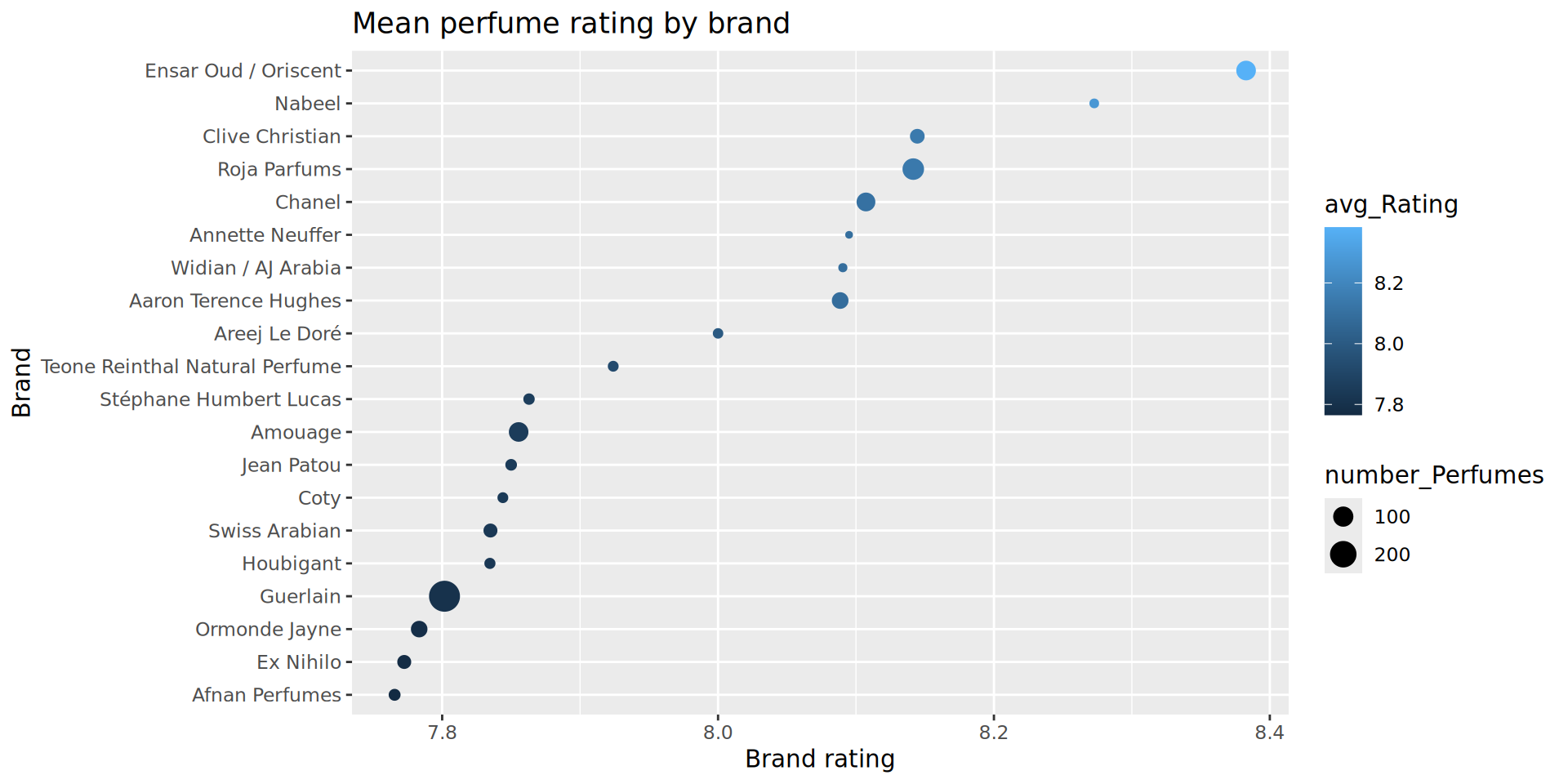
Refining the plot
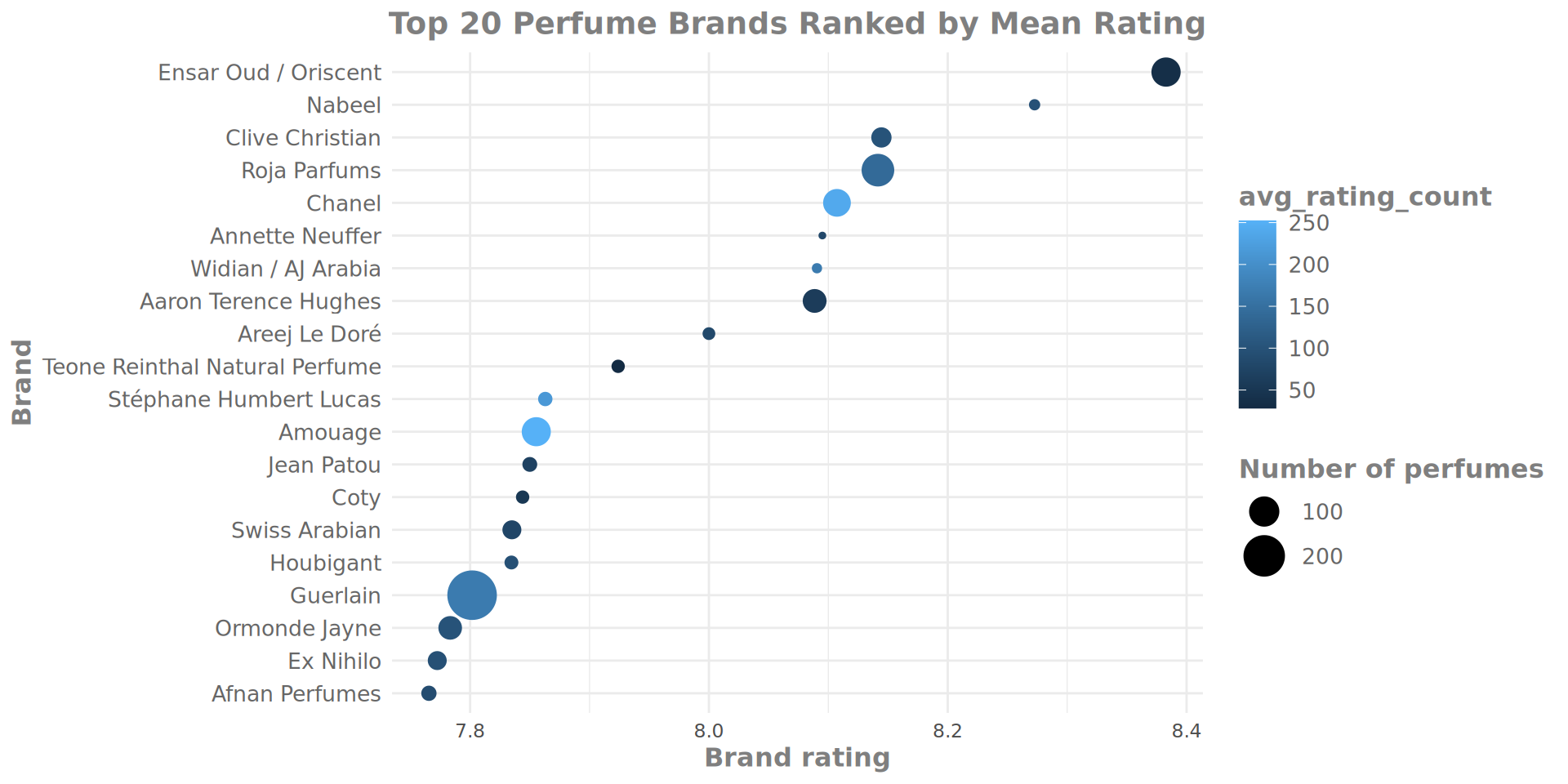
Currently, colour is not being fully utilised - it is mapped to the Brand average rating, which is already shown via the x axis. Thinking back to earlier themes on number of ratings and number of perfumes per brand, we could use colour to represent some of our data around the rating_Count variable.
To do this, we will re-build the brand_rating_data object with an additional variable calculated by the summarize() function, calling it avg_rating_count and then map it to colour.
Refined
Summary
ggplot has fine control over all elements of the plot window.
Combine geoms, separate data with facet(), join figures with patchwork.
ggplot facilitates exploratory data analysis with quick plots.
dplyr verbs can manage majority of data wrangling.
Plot, transform, plot with refined features.
Before we stop
Open online books
H. Wickham R for data science
H. Wickham ggplot book
Jenny Bryan Happy git with R
Next courses
Linear algebra and statistics with R - 14-16 January 2026
Data transformation with tidyverse - 6 -8 February 2026
Data visualisation - February - May (weekly) 2026
Advanced R - June 2026

